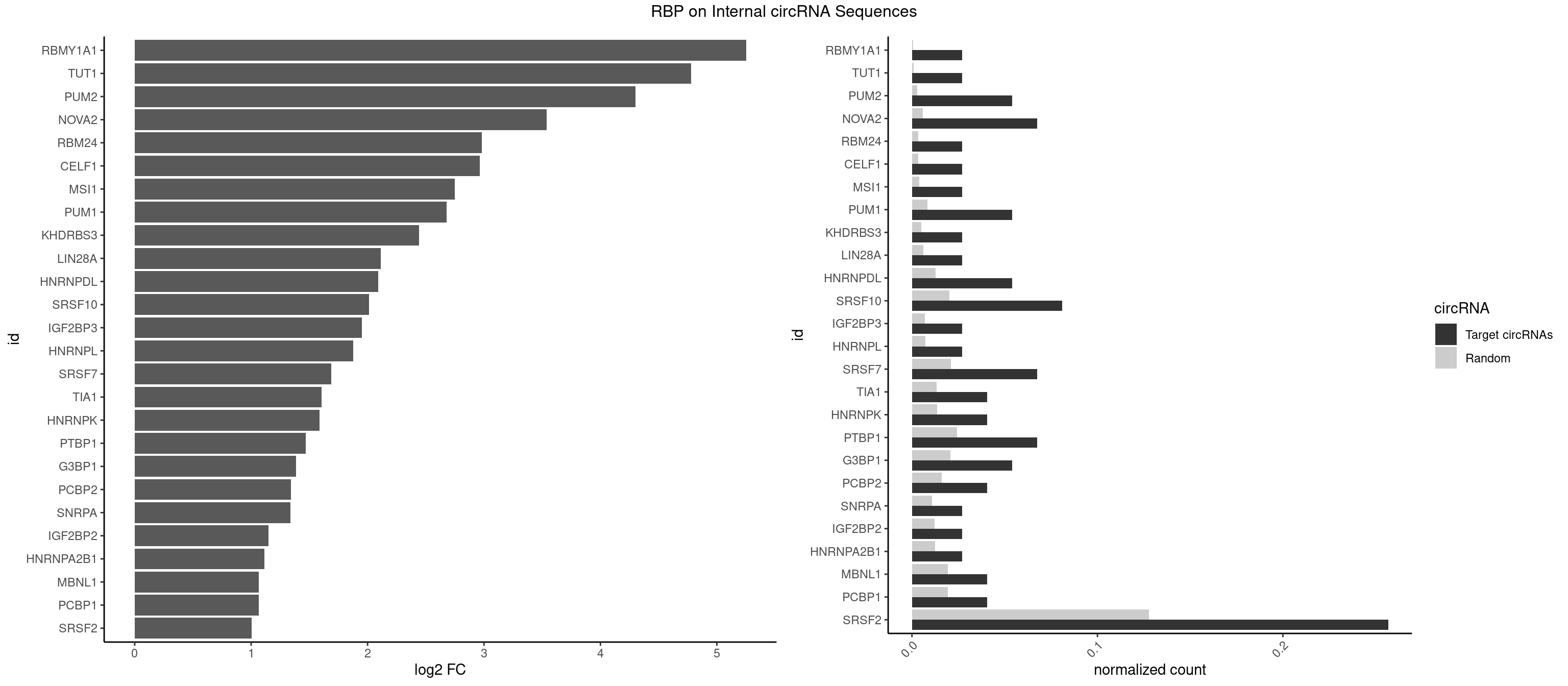
| RBMY1A1 |
1 |
489 |
0.02702703 |
0.0007101099 |
5.250217 |
ACAAGA |
ACAAGA,CAAGAC |
| TUT1 |
1 |
678 |
0.02702703 |
0.0009840095 |
4.779587 |
AAAUAC |
AAAUAC,AAUACU,AGAUAC,CAAUAC,CGAUAC,GAUACU |
| PUM2 |
3 |
1890 |
0.05405405 |
0.0027404447 |
4.301921 |
GUAAAU,UAAAUA,UGUAAA |
GUAAAU,GUACAU,GUAGAU,GUAUAU,UAAAUA,UACAUA,UACAUC,UAGAUA,UAUAUA,UGUAAA,UGUACA,UGUAGA,UGUAUA |
| NOVA2 |
4 |
4013 |
0.06756757 |
0.0058171047 |
3.537958 |
AGAUCA,CUAGAU,GAUCAC,UAGAUC |
AACACC,ACCACC,AGACAU,AGAUCA,AGCACC,AGGCAU,AGUCAU,AUCAAC,AUCACC,AUCAUC,CCUAGA,CUAGAU,GAGACA,GAGGCA,GAGUCA,GAUCAC,GGGGGG,UAGAUC,UUUUUU |
| RBM24 |
1 |
2357 |
0.02702703 |
0.0034172229 |
2.983507 |
GAGUGG |
AGAGUG,AGUGUG,GAGUGA,GAGUGG,GAGUGU,GUGUGA,GUGUGG,GUGUGU,UGAGUG,UGUGUG |
| CELF1 |
1 |
2391 |
0.02702703 |
0.0034664959 |
2.962853 |
CUGUCU |
CUGUCU,GUGUGU,GUGUUG,GUUGUG,GUUUGU,UGUCUG,UGUGUG,UGUGUU,UGUUGU,UGUUUG,UUGUGU |
| MSI1 |
1 |
2770 |
0.02702703 |
0.0040157442 |
2.750664 |
UAGGAG |
AGGAAG,AGGAGG,AGGUAG,AGGUGG,AGUAAG,AGUAGG,AGUUAG,AGUUGG,UAGGAA,UAGGAG,UAGGUA,UAGGUG,UAGUAA,UAGUAG,UAGUUA,UAGUUG |
| PUM1 |
3 |
5823 |
0.05405405 |
0.0084401638 |
2.679060 |
GUAAAU,UAAAUA,UGUAAA |
AAUAUU,AAUGUU,AAUUGU,ACAUAA,AGAAUU,AGAUAA,AUUGUA,CAGAAU,CCAGAA,CUUGUA,GAAUUG,GUAAAU,GUAAUA,GUACAU,GUAGAU,GUAUAU,GUCCAG,UAAAUA,UAAUAU,UAAUGU,UACAUA,UACAUC,UAGAUA,UAUAUA,UGUAAA,UGUAAU,UGUACA,UGUAGA,UGUAUA,UGUCCA,UUAAUG,UUGUAC,UUGUAG,UUUAAU |
| KHDRBS3 |
1 |
3429 |
0.02702703 |
0.0049707696 |
2.442862 |
UAAAUA |
AAAUAA,AAUAAA,AGAUAA,AUAAAA,AUAAAC,AUAAAG,AUUAAA,GAUAAA,UAAAAC,UAAAUA,UUAAAC,UUUUUU |
| LIN28A |
1 |
4315 |
0.02702703 |
0.0062547643 |
2.111375 |
AGGAGU |
AGGAGA,AGGAGU,CGGAGA,CGGAGG,CGGAGU,GGAGAA,GGAGAU,GGAGGA,GGAGGG,GGAGUA,UGGAGA,UGGAGG,UGGAGU |
| HNRNPDL |
3 |
8759 |
0.05405405 |
0.0126950266 |
2.090139 |
AAUACC,CUAGAU,CUAGGA |
AACAGC,AACCUU,AACUAA,AAUACC,AAUUUA,ACACCA,ACAGCA,ACAUUA,ACCACG,ACCAGA,ACCUUG,ACUAAC,ACUAAG,ACUAGA,ACUAGC,ACUAGG,ACUGCA,ACUUUA,AGAUAU,AGUAGC,AGUAGG,AUACCA,AUCUGA,AUGCGC,AUUAGC,AUUAGG,CACCAG,CACGCA,CACUAG,CAUUAG,CCACGC,CCAGAC,CCUUGC,CCUUUA,CGAGCA,CUAACA,CUAACU,CUAAGC,CUAAGU,CUAGAG,CUAGAU,CUAGCA,CUAGCC,CUAGCG,CUAGCU,CUAGGA,CUAGGC,CUAGUA,CUUGCC,CUUUAA,CUUUAG,GAACUA,GACUAG,GAGUAG,GAUUAG,GCACUA,GCGAGC,GCUAGU,GGACUA,GGAGUA,GGAUUA,GUAGCC,GUAGCU,UAACAG,UAAGUA,UAGGCA,UCUGAC,UGCGCA,UGUCGC,UUAGCC,UUAGGC,UUUAGG |
| SRSF10 |
5 |
13860 |
0.08108108 |
0.0200874160 |
2.013073 |
AAGAGA,AGAGAA,AGAGAG,GAGAAG,GAGAGA |
AAAAGA,AAAGAA,AAAGAC,AAAGAG,AAAGGA,AAAGGG,AAGAAA,AAGAAG,AAGACA,AAGAGA,AAGAGG,AAGGAA,AAGGAG,AAGGGA,AAGGGG,ACAAAG,AGACAA,AGAGAA,AGAGAC,AGAGAG,AGAGGA,AGAGGG,CAAAGA,GAAAGA,GACAAA,GAGAAA,GAGAAC,GAGAAG,GAGACA,GAGACC,GAGACG,GAGAGA,GAGAGC,GAGAGG,GAGGAA,GAGGAG,GAGGGA,GAGGGG |
| IGF2BP3 |
1 |
4815 |
0.02702703 |
0.0069793662 |
1.953235 |
AAAUAC |
AAAAAC,AAAACA,AAAAUC,AAACAC,AAACUC,AAAUAC,AAAUCA,AAAUUC,AACACA,AACUCA,AAUACA,AAUUCA,ACAAAC,ACAAUC,ACACAC,ACACUC,ACAUAC,ACAUUC,CAAACA,CAAUCA,CACACA,CACUCA,CAUACA,CAUUCA |
| HNRNPL |
1 |
5085 |
0.02702703 |
0.0073706513 |
1.874539 |
AAAUAC |
AAACAA,AAACAC,AAAUAA,AAAUAC,AACAAA,AACACA,AAUAAA,AAUACA,ACACAA,ACACAC,ACACCA,ACACGA,ACAUAA,ACAUAC,ACCACA,CACAAA,CACAAC,CACAAG,CACACA,CACACC,CACCAC,CACGAA,CACGAC,CACGAG,CAUAAA,CAUACA |
| SRSF7 |
4 |
14481 |
0.06756757 |
0.0209873716 |
1.686809 |
AGAGAA,AGAGAG,AGAUCA,GAGAGA |
AAAGGA,AAGAAG,AAGGAC,AAGGCG,AAUGAU,ACGAAU,ACGACG,ACGAGA,ACGAUA,ACGAUU,ACUACG,ACUAGA,AGAAGA,AGACGA,AGACUA,AGAGAA,AGAGAC,AGAGAG,AGAGAU,AGAGGA,AGAUCA,AGAUCU,AGGAAG,AGGACA,AGGCGA,AUAGAA,AUAGAC,AUUGAC,AUUGAU,CAGAGA,CCGAGA,CGAAUG,CGACGA,CGAGAC,CGAGAG,CGAGAU,CGAUAG,CGAUUG,CUACGA,CUAGAG,CUCUUC,CUGAGA,CUUCAC,GAAGAA,GAAGGC,GAAUGA,GACAAA,GACGAC,GACGAG,GACGAU,GACUAC,GAGACU,GAGAGA,GAGAUC,GAGGAA,GAUAGA,GAUUGA,GGAAGG,GGACAA,GGACGA,UACGAC,UACGAU,UAGAGA,UCAACA,UCGAGA,UCGAUU,UCUCCG,UCUUCA,UGAGAG,UGGACA |
| TIA1 |
2 |
9206 |
0.04054054 |
0.0133428208 |
1.603302 |
CUUUUC,UUUUCU |
AUUUUC,AUUUUG,AUUUUU,CCUUUC,CUCCUU,CUUUUA,CUUUUC,CUUUUG,CUUUUU,GUUUUA,GUUUUC,GUUUUG,GUUUUU,UAUUUU,UCCUUU,UCUCCU,UUAUUU,UUCUCC,UUUAUU,UUUUAU,UUUUCG,UUUUCU,UUUUGG,UUUUGU,UUUUUA,UUUUUC,UUUUUG,UUUUUU |
| HNRNPK |
2 |
9304 |
0.04054054 |
0.0134848428 |
1.588027 |
CCCCAC,UCCCCA |
AAAAAA,ACCCAA,ACCCAU,ACCCCA,ACCCCC,ACCCGA,ACCCGC,ACCCGU,ACCCUU,CAAACC,CAACCC,CAUACC,CAUCCC,CCAAAC,CCAACC,CCAUAC,CCAUCC,CCCCAA,CCCCAC,CCCCAU,CCCCCA,CCCCCC,CCCCCU,CCCCUA,CCCCUU,GCCCAA,GCCCAC,GCCCAG,GCCCCC,GCCCCU,GGGGGG,UCCCAA,UCCCAC,UCCCAU,UCCCCA,UCCCCC,UCCCCU,UCCCGA,UCCCUA,UCCCUU,UUUUUU |
| PTBP1 |
4 |
16854 |
0.06756757 |
0.0244263326 |
1.467894 |
CUCCCC,CUUUUC,GCUCCC,UUUUCU |
ACUUUC,ACUUUU,AGCUGU,AUCUUC,AUUUUC,AUUUUU,CAUCUU,CCAUCU,CCCCCC,CCUCUU,CCUUCC,CCUUUC,CCUUUU,CUAUCU,CUCCAU,CUCCCC,CUCUCU,CUCUUA,CUCUUC,CUCUUG,CUCUUU,CUUCCU,CUUCUC,CUUCUU,CUUUCC,CUUUCU,CUUUUC,CUUUUU,GCUCCC,GCUGUG,GGCUCC,GGGGGG,GUCUUA,GUCUUU,UACUUU,UAGCUG,UAUUUU,UCCAUC,UCCCCC,UCCUCU,UCUAUC,UCUCUC,UCUCUU,UCUUCU,UCUUUC,UCUUUU,UUACUU,UUAUUU,UUCCCC,UUCCUU,UUCUCU,UUCUUC,UUCUUG,UUCUUU,UUUCCC,UUUCUU,UUUUCC,UUUUCU,UUUUUC,UUUUUU |
| G3BP1 |
3 |
14282 |
0.05405405 |
0.0206989801 |
1.384843 |
CCCACG,CCCCAC,CCUAGG |
ACACGC,ACAGGC,ACCCAC,ACCCAU,ACCCCC,ACCCCU,ACCCGC,ACCGGC,ACGCAC,ACGCAG,ACGCCC,ACGCCG,AGGCAC,AGGCAG,AGGCCC,AGGCCG,AUACGC,AUAGGC,AUCCGC,AUCGGC,CACACG,CACAGG,CACCCG,CACCGG,CACGCA,CACGCC,CAGGCA,CAGGCC,CAUACG,CAUAGG,CAUCCG,CAUCGG,CCACAC,CCACAG,CCACCC,CCACCG,CCACGC,CCAGGC,CCAUAC,CCAUAG,CCAUCC,CCAUCG,CCCACA,CCCACC,CCCACG,CCCAGG,CCCAUA,CCCAUC,CCCCAC,CCCCAG,CCCCCA,CCCCCC,CCCCCG,CCCCGC,CCCCGG,CCCCUA,CCCCUC,CCCGCA,CCCGCC,CCCGGC,CCCUAC,CCCUAG,CCCUCC,CCCUCG,CCGCAC,CCGCAG,CCGCCC,CCGCCG,CCGGCA,CCGGCC,CCUACG,CCUAGG,CCUCCG,CCUCGG,CGGCAC,CGGCAG,CGGCCC,CGGCCG,CUACGC,CUAGGC,CUCCGC,CUCGGC,UACGCA,UACGCC,UAGGCA,UAGGCC,UCCGCA,UCCGCC,UCGGCA,UCGGCC |
| PCBP2 |
2 |
11045 |
0.04054054 |
0.0160079069 |
1.340581 |
CUCCCC,UCCCCA |
AAACCA,AAAUUA,AAAUUC,AACCAA,AACCCU,AAUUAA,AAUUAC,AAUUCA,AAUUCC,ACAUUA,ACCAAA,ACCCUA,ACCUUA,AUUAAA,AUUACC,AUUCAC,AUUCCA,AUUCCC,CAAUUA,CAUUAA,CCAUUA,CCAUUC,CCCCCA,CCCCCC,CCCCCU,CCCUAA,CCCUCC,CCCUUA,CCCUUC,CCUCCA,CCUCCC,CCUCCU,CCUUAA,CCUUCC,CUAACC,CUCCCA,CUCCCC,CUCCCU,CUUAAA,CUUCCA,CUUCCC,CUUCCU,GGGGGG,UAACCC,UCCCCA,UUAGAG,UUAGGA,UUAGGG,UUUUUU |
| SNRPA |
1 |
7380 |
0.02702703 |
0.0106965744 |
1.337254 |
AUACCU |
ACCUGC,AGCAGU,AGGAGA,AGUAGG,AGUAGU,AUACCU,AUGCAC,AUGCUG,AUUCCU,AUUGCA,CAGUAG,CCUGCU,CUGCUA,GAGCAG,GAUACC,GAUUCC,GCAGUA,GGAGAU,GGGUAU,GGUAUG,GUAGGC,GUAGGG,GUAGUC,GUAGUG,GUAUGC,GUUACC,GUUUCC,UACCUG,UAUGCU,UCCUGC,UGCACA,UGCACC,UGCACG,UUACCU,UUCCUG,UUGCAC,UUUCCU |
| IGF2BP2 |
1 |
8408 |
0.02702703 |
0.0121863560 |
1.149136 |
AAAUAC |
AAAAAC,AAAACA,AAAAUC,AAACAC,AAACUC,AAAUAC,AAAUCA,AAAUUC,AACACA,AACUCA,AAUACA,AAUUCA,ACAAAC,ACAAUC,ACACAC,ACACUC,ACAUAC,ACAUUC,CAAAAC,CAAACA,CAAAUC,CAACAC,CAACUC,CAAUAC,CAAUCA,CAAUUC,CACACA,CACUCA,CAUACA,CAUUCA,CCAAAC,CCAAUC,CCACAC,CCACUC,CCAUAC,CCAUUC,GAAAAC,GAAAUC,GAACAC,GAACUC,GAAUAC,GAAUUC,GCAAAC,GCAAUC,GCACAC,GCACUC,GCAUAC,GCAUUC |
| HNRNPA2B1 |
1 |
8607 |
0.02702703 |
0.0124747476 |
1.115392 |
AGAAGC |
AAGAAG,AAGGAA,AAGGAG,AAGGGG,AAUUUA,ACUAGA,AGAAGC,AGACUA,AGAUAU,AGGAAC,AGGAGC,AGGGGC,AGUAGG,AUAGCA,AUAGGG,AUUUAA,CAAGAA,CAAGGA,CAAGGG,CCAAGA,CCAAGG,CGAAGG,CUAGAC,GAAGCC,GAAGGA,GACUAG,GCCAAG,GCGAAG,GGAACC,GGAGCC,GGGGCC,GUAGGG,UAGACA,UAGACU,UAGGGA,UAGGGU,UUAGGG,UUUAUA |
| MBNL1 |
2 |
13372 |
0.04054054 |
0.0193802045 |
1.064782 |
GCUUUU,GGCUUU |
ACGCUA,ACGCUC,ACGCUG,ACGCUU,AUGCCA,AUGCCC,AUGCCG,AUGCUC,AUGCUU,CCCGCU,CCGCUA,CCGCUC,CCGCUG,CCGCUU,CCUGCU,CGCUGC,CGCUGU,CGCUUC,CGCUUG,CGCUUU,CUCGCU,CUGCCA,CUGCCC,CUGCCG,CUGCCU,CUGCGG,CUGCUA,CUGCUC,CUGCUG,CUGCUU,CUUGCU,CUUGUG,GCGCUC,GCGCUG,GCGCUU,GCUGCG,GCUGCU,GCUUGC,GCUUGU,GCUUUU,GGCUUU,GUCUCG,GUGCUA,GUGCUC,GUGCUG,GUGCUU,UCCGCU,UCGCUA,UCGCUC,UCGCUG,UCGCUU,UCUCGC,UCUGCU,UGCGGC,UGCUGC,UGCUGU,UGCUUC,UGCUUU,UGUCUC,UUCGCU,UUGCCA,UUGCCC,UUGCCG,UUGCCU,UUGCUA,UUGCUC,UUGCUG,UUGCUU,UUGUGC,UUUGCU |
| PCBP1 |
2 |
13384 |
0.04054054 |
0.0193975949 |
1.063488 |
CCCCAC,UCCCCA |
AAAAAA,AAACCA,AAAUUA,AAAUUC,AACCAA,AAUUAA,AAUUAC,AAUUCA,AAUUCC,ACAUUA,ACCAAA,ACCCUC,ACCUUA,AUUAAA,AUUACC,AUUCAC,AUUCCA,AUUCCC,CAAACC,CAACCC,CAAUCC,CAAUUA,CACCCU,CAUACC,CAUCCC,CAUUAA,CAUUCC,CCAAAC,CCAACC,CCAAUC,CCACCC,CCAUAC,CCAUCC,CCAUUA,CCAUUC,CCCACC,CCCCAA,CCCCAC,CCCCCC,CCCUCC,CCCUCU,CCCUUA,CCCUUC,CCUAAC,CCUACC,CCUAUC,CCUCCC,CCUCUU,CCUUAA,CCUUAC,CCUUCC,CCUUUC,CUAACC,CUACCC,CUAUCC,CUUAAA,CUUACC,CUUCCC,CUUUCC,GGGGGG,UCCCCA,UUAGAG,UUAGGA,UUAGGG,UUUUUU |
| SRSF2 |
18 |
88240 |
0.25675676 |
0.1278792060 |
1.005621 |
AAGAGA,AAUACC,AGAAGC,AGAGAA,AGCUCC,AGGAGU,CGUGUA,GACUGU,GAGAAG,GAGAGA,GAUCAC,GCUCCC,GGACUG,GGAGUG,GGUUCC,GUGUAA,GUUCCG,UCCCCA |
AAAAGA,AAAGAG,AAAGGA,AACCAG,AACGAG,AAGAAG,AAGAGA,AAGCAG,AAGCGG,AAGCUG,AAGGAG,AAGGCG,AAGUAC,AAUACC,AAUCCC,AAUGCC,AAUGCU,ACCACC,ACCACU,ACCAGC,ACCAGU,ACCCCC,ACCCCU,ACCCGC,ACCCGG,ACCGGU,ACCGUA,ACCUCC,ACCUGC,ACGAGA,ACGAGG,ACGCUG,ACGGAG,ACGUGU,ACUACC,ACUACU,ACUAGA,ACUCAA,ACUCCC,ACUCCU,ACUGUA,AGAAGA,AGAAGC,AGAAUA,AGAAUG,AGACUA,AGAGAA,AGAGAU,AGAGGA,AGAGGU,AGAGUA,AGAGUG,AGAGUU,AGAUAA,AGAUCC,AGAUGC,AGCACU,AGCAGA,AGCAGU,AGCCCC,AGCCGA,AGCCUC,AGCGGA,AGCGUG,AGCUCC,AGCUGU,AGGAAG,AGGAGA,AGGAGC,AGGAGG,AGGAGU,AGGCAG,AGGCGA,AGGCGU,AGGCUG,AGGGUA,AGUACG,AGUAGA,AGUAGG,AGUAGU,AGUAUC,AGUCGA,AGUGAC,AGUGGU,AGUGUU,AGUUCC,AUACCG,AUCACC,AUCACU,AUCCCC,AUCCCG,AUCCCU,AUCCGG,AUCGAG,AUCGCU,AUGAAU,AUGAUG,AUGCCG,AUGCGU,AUGCUG,AUGGAG,AUUACC,AUUACU,AUUAGU,AUUCCA,AUUCCC,AUUCCU,AUUGAU,CACCUC,CAGAGA,CAGAGG,CAGAGU,CAGCUA,CAGUAG,CAGUCC,CAGUGG,CAGUGU,CCACCA,CCACCG,CCACUA,CCACUG,CCAGAA,CCAGAU,CCAGCC,CCAGCU,CCAGGG,CCAGGU,CCAGUC,CCAGUG,CCAGUU,CCCACA,CCCCCA,CCCCCG,CCCCUA,CCCCUG,CCCGCA,CCCGCU,CCCGUG,CCCUUG,CCGAGA,CCGCAG,CCGCUA,CCGGAU,CCGGUG,CCUCAG,CCUCCG,CCUGCG,CCUGCU,CCUGUU,CGAACG,CGAGAA,CGAGAG,CGAGAU,CGAGCA,CGAGGA,CGAGUA,CGAGUG,CGAGUU,CGCAGG,CGCAGU,CGCCCU,CGCUGC,CGGAGA,CGGAGU,CGUAAG,CGUGAU,CGUGCG,CGUGUA,CGUGUU,CUAACG,CUACCA,CUACCG,CUACUA,CUACUG,CUAGAA,CUAGAC,CUCAAG,CUCCAA,CUCCCA,CUCCCG,CUCCUA,CUCCUG,CUCGAG,CUCGUG,CUGAUG,CUGCAA,CUGCUA,CUGUUA,GAAAGG,GAACCA,GAACGA,GAAGAA,GAAGCG,GAAGGC,GAAUAC,GAAUCC,GAAUGC,GACCAC,GACCCC,GACCCG,GACCGG,GACCGU,GACCUG,GACGCU,GACGGA,GACUAC,GACUAG,GACUCA,GACUCC,GACUGU,GAGAAG,GAGAAU,GAGAGA,GAGAGG,GAGAGU,GAGAUA,GAGAUG,GAGCAC,GAGCAG,GAGCGU,GAGCUG,GAGGAA,GAGGAC,GAGGAG,GAGUAU,GAGUGA,GAUCAC,GAUCCC,GAUCCG,GAUCGA,GAUCGC,GAUGAU,GAUGCG,GAUGCU,GAUGGA,GAUUAC,GAUUAG,GAUUCC,GAUUGA,GCACUG,GCAGAG,GCAGGG,GCAGGU,GCAGUA,GCAGUG,GCCAAA,GCCACC,GCCACU,GCCCAC,GCCCCC,GCCCCU,GCCCUU,GCCGAG,GCCGCA,GCCGCC,GCCGUU,GCCUCA,GCCUCC,GCGAGC,GCGAGU,GCGCAG,GCGGAG,GCGGUC,GCGUGU,GCUACC,GCUACU,GCUCCA,GCUCCC,GCUCCU,GCUCGA,GCUCGU,GCUGAU,GCUGCA,GCUGCU,GCUGUU,GGAACC,GGAAGG,GGAAUG,GGACCG,GGACGC,GGACUG,GGAGAA,GGAGAG,GGAGAU,GGAGCC,GGAGCG,GGAGGA,GGAGUA,GGAGUG,GGAGUU,GGAUCC,GGAUGG,GGCAGU,GGCCAA,GGCCAC,GGCCCA,GGCCCC,GGCCGC,GGCCUC,GGCGAG,GGCGUG,GGCUAC,GGCUCC,GGCUCG,GGCUGA,GGCUGC,GGGAAU,GGGAGC,GGGCAG,GGGUAA,GGGUAC,GGGUAG,GGGUAU,GGGUGA,GGUAAG,GGUACG,GGUAGG,GGUAUG,GGUCAC,GGUCAG,GGUCCC,GGUCGC,GGUGAG,GGUGAU,GGUUAA,GGUUAC,GGUUCC,GGUUGG,GGUUGU,GUAACC,GUAAGC,GUAAGU,GUACGA,GUACGC,GUAGGC,GUAGGG,GUAGUC,GUAUCC,GUAUGC,GUCACC,GUCACU,GUCAGU,GUCCCC,GUCCCU,GUCCUC,GUCGAU,GUCGCC,GUCUAA,GUGAGC,GUGAUG,GUGCAG,GUGGUC,GUGUAA,GUUAAU,GUUACC,GUUACG,GUUACU,GUUCCA,GUUCCC,GUUCCG,GUUCCU,GUUCGA,GUUCGU,GUUCUG,GUUGGC,GUUGUU,GUUUCG,UAACCA,UAAGCU,UAAGUA,UACCAG,UACGAG,UACGCU,UACGUG,UAGACU,UAGGCU,UAUCCC,UAUGCC,UAUGCU,UCACCA,UCACCG,UCACUA,UCACUG,UCAGUG,UCCAGA,UCCAGC,UCCAGG,UCCAGU,UCCCCA,UCCCCG,UCCCCU,UCCCGC,UCCCGU,UCCCUA,UCCCUG,UCCGGA,UCCUCA,UCCUCC,UCCUGC,UCCUGU,UCGAGA,UCGAGU,UCGCCG,UCGCUG,UCGUAA,UCGUGC,UCUAAC,UCUGUA,UGAAUG,UGAGCU,UGAUCG,UGAUGC,UGAUGG,UGCAGA,UGCAGU,UGCCGC,UGCCGU,UGCGGU,UGCUGU,UGGAGA,UGGAGG,UGGAGU,UGGCAG,UGUUAC,UGUUCC,UGUUCG,UUAAUG,UUACCA,UUACCG,UUACGU,UUACUA,UUACUG,UUAGUG,UUCCAG,UUCCCA,UUCCCG,UUCCCU,UUCCGG,UUCCUA,UUCCUG,UUCGAG,UUCGUG,UUCUGU,UUGAUG,UUGGCG,UUGUUG,UUUCGA |