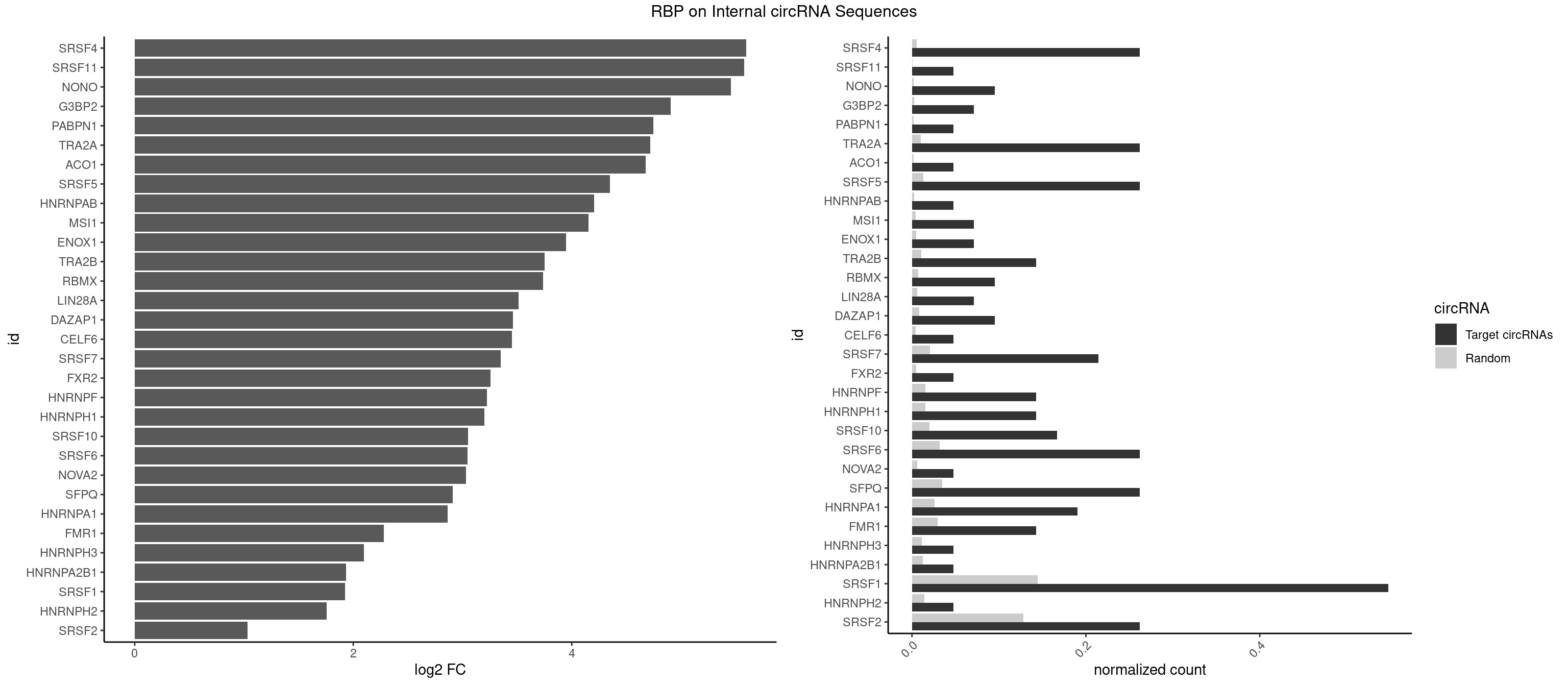
| SRSF4 |
10 |
3740 |
0.26190476 |
0.0054214720 |
5.594214 |
AAGAAG,AGAAGA,AGGAAG,GAAGAA,GAGGAA,GGAAGA |
AAGAAG,AGAAGA,AGGAAG,GAAGAA,GAGGAA,GGAAGA |
| SRSF11 |
1 |
688 |
0.04761905 |
0.0009985015 |
5.575630 |
AAGAAG |
AAGAAG |
| NONO |
3 |
1498 |
0.09523810 |
0.0021723567 |
5.454206 |
AGAGGA,GAGGAA |
AGAGGA,AGGAAC,GAGAGG,GAGGAA |
| G3BP2 |
2 |
1644 |
0.07142857 |
0.0023839405 |
4.905081 |
AGGAUG,GGAUGA |
AGGAUA,AGGAUG,AGGAUU,GGAUAA,GGAUAG,GGAUGA,GGAUGG,GGAUUA,GGAUUG |
| PABPN1 |
1 |
1222 |
0.04761905 |
0.0017723764 |
4.747782 |
AGAAGA |
AAAAGA,AGAAGA |
| TRA2A |
10 |
6871 |
0.26190476 |
0.0099589296 |
4.716908 |
AAGAAG,AAGAGG,AGAAGA,AGAGGA,AGGAAG,GAAGAA,GAAGAG,GAGGAA |
AAAGAA,AAGAAA,AAGAAG,AAGAGG,AGAAAG,AGAAGA,AGAGGA,AGGAAG,GAAAGA,GAAGAA,GAAGAG,GAGGAA |
| ACO1 |
1 |
1283 |
0.04761905 |
0.0018607779 |
4.677561 |
CAGUGA |
CAGUGA,CAGUGC,CAGUGG,CAGUGU |
| SRSF5 |
10 |
8869 |
0.26190476 |
0.0128544391 |
4.348704 |
AAGAAG,AGAAGA,AGGAAG,GAAGAA,GAGGAA,GGAAGA |
AACAGC,AAGAAG,AGAAGA,AGGAAG,AUAAAG,CAACAG,CACAGA,CACAGC,CACAUA,CACAUC,CACGGA,CACGGC,CACGUA,CACGUC,CCGCAG,CGCAGA,CGCAGC,CGCAGG,CGCAUA,CGCAUC,CGCGGA,CGCGGC,CGCGUA,CGCGUC,GAAGAA,GAGGAA,GGAAGA,UAAAGG,UACAGA,UACAGC,UACAUA,UACAUC,UACGGA,UACGGC,UACGUA,UACGUC,UGCAGA,UGCAGC,UGCAUA,UGCAUC,UGCGGA,UGCGGC,UGCGUA,UGCGUC |
| HNRNPAB |
1 |
1782 |
0.04761905 |
0.0025839306 |
4.203900 |
AAGACA |
AAAGAC,AAGACA,ACAAAG,AGACAA,AUAGCA,CAAAGA,GACAAA |
| MSI1 |
2 |
2770 |
0.07142857 |
0.0040157442 |
4.152762 |
AGGAAG |
AGGAAG,AGGAGG,AGGUAG,AGGUGG,AGUAAG,AGUAGG,AGUUAG,AGUUGG,UAGGAA,UAGGAG,UAGGUA,UAGGUG,UAGUAA,UAGUAG,UAGUUA,UAGUUG |
| ENOX1 |
2 |
3195 |
0.07142857 |
0.0046316558 |
3.946901 |
AAGACA,AGACAG |
AAGACA,AAUACA,AGACAG,AGGACA,AGUACA,AUACAG,CAGACA,CAUACA,CGGACA,CGUACA,GGACAG,GUACAG,UAGACA,UAUACA,UGGACA,UGUACA |
| TRA2B |
5 |
7329 |
0.14285714 |
0.0106226650 |
3.749356 |
AAGAAG,AGAAGA,AGGAAG,GAAGAA |
AAAGAA,AAGAAC,AAGAAG,AAGAAU,AAGGAA,AAUAAG,AGAAGA,AGAAGG,AGGAAA,AGGAAG,AUAAGA,GAAAGA,GAAGAA,GAAGGA,GAAUUA,GGAAGG,UAAGAA |
| RBMX |
3 |
4925 |
0.09523810 |
0.0071387787 |
3.737790 |
AAGAAG,AGGAAG |
AAGAAG,AAGGAA,AAGUAA,AAGUGU,ACCAAA,AGAAGG,AGGAAG,AGUAAC,AGUGUU,AUCAAA,AUCCCA,AUCCCC,GAAGGA,GGAAGG,GUAACA,UAACAA,UAAGAC,UCAAAA |
| LIN28A |
2 |
4315 |
0.07142857 |
0.0062547643 |
3.513474 |
CGGAGG,GGAGGA |
AGGAGA,AGGAGU,CGGAGA,CGGAGG,CGGAGU,GGAGAA,GGAGAU,GGAGGA,GGAGGG,GGAGUA,UGGAGA,UGGAGG,UGGAGU |
| DAZAP1 |
3 |
5964 |
0.09523810 |
0.0086445016 |
3.461684 |
AGGAAG,AGGAUG |
AAAAAA,AAUUUA,AGAUAU,AGGAAA,AGGAAG,AGGAUA,AGGAUG,AGGUAA,AGGUAG,AGGUUA,AGGUUG,AGUAAA,AGUAAG,AGUAGG,AGUAUA,AGUAUG,AGUUAA,AGUUAG,AGUUUA,AGUUUG,GGGGGG,GUAACG,UAGGAA,UAGGAU,UAGGUA,UAGGUU,UAGUAA,UAGUAU,UAGUUA,UAGUUU,UUUUUU |
| CELF6 |
1 |
2997 |
0.04761905 |
0.0043447134 |
3.454206 |
GUGAGG |
GUGAGG,GUGAUG,GUGGGG,GUGGUG,GUGUGG,GUGUUG,UGUGAG,UGUGAU,UGUGGG,UGUGGU,UGUGUG,UGUGUU |
| SRSF7 |
8 |
14481 |
0.21428571 |
0.0209873716 |
3.351942 |
AAGAAG,AGAAGA,AGAGGA,AGGAAG,GAAGAA,GAGGAA |
AAAGGA,AAGAAG,AAGGAC,AAGGCG,AAUGAU,ACGAAU,ACGACG,ACGAGA,ACGAUA,ACGAUU,ACUACG,ACUAGA,AGAAGA,AGACGA,AGACUA,AGAGAA,AGAGAC,AGAGAG,AGAGAU,AGAGGA,AGAUCA,AGAUCU,AGGAAG,AGGACA,AGGCGA,AUAGAA,AUAGAC,AUUGAC,AUUGAU,CAGAGA,CCGAGA,CGAAUG,CGACGA,CGAGAC,CGAGAG,CGAGAU,CGAUAG,CGAUUG,CUACGA,CUAGAG,CUCUUC,CUGAGA,CUUCAC,GAAGAA,GAAGGC,GAAUGA,GACAAA,GACGAC,GACGAG,GACGAU,GACUAC,GAGACU,GAGAGA,GAGAUC,GAGGAA,GAUAGA,GAUUGA,GGAAGG,GGACAA,GGACGA,UACGAC,UACGAU,UAGAGA,UCAACA,UCGAGA,UCGAUU,UCUCCG,UCUUCA,UGAGAG,UGGACA |
| FXR2 |
1 |
3434 |
0.04761905 |
0.0049780156 |
3.257896 |
AGACAG |
AGACAA,AGACAG,AGACGA,AGACGG,GACAAA,GACAAG,GACAGA,GACAGG,GACGAA,GACGAG,GACGGA,GACGGG,GGACAA,GGACAG,GGACGA,GGACGG,UGACAA,UGACAG,UGACGA,UGACGG |
| HNRNPF |
5 |
10561 |
0.14285714 |
0.0153064921 |
3.222358 |
AGGAAG,GAGGAA,GGAGGA |
AAGGCG,AAGGGA,AAGGGG,AAGGUG,AAUGUG,AGGAAG,AGGCGA,AGGGAA,AGGGAU,AGGGGA,AUGGGA,AUGGGG,AUGUGG,CGAUGG,CGGGAU,CGGGGG,CUGGGG,GAAGGG,GAAGGU,GAAUGU,GAGGAA,GAGGGG,GAUGGG,GGAAGG,GGAAUG,GGAGGA,GGAGGG,GGAUGG,GGCGAA,GGGAAG,GGGAAU,GGGAGG,GGGAUG,GGGCUG,GGGGAA,GGGGAG,GGGGCU,GGGGGC,GGGGGG,GGGGUA,UGGGAA,UGGGGU,UGUGGG |
| HNRNPH1 |
5 |
10717 |
0.14285714 |
0.0155325680 |
3.201205 |
AGGAAG,GAGGAA,GGAGGA |
AAGGCG,AAGGGA,AAGGGG,AAGGUG,AAUGUG,AGGAAG,AGGCGA,AGGGAA,AGGGGA,AUCGGG,AUGUGG,AUUGGG,CAGGAC,CGAGGG,CGGGCG,CGGGGG,CUGGGG,GAAGGG,GAAGGU,GAAUGU,GAGGAA,GAGGGG,GGAAGG,GGAAUG,GGAGGA,GGAGGG,GGCGAA,GGGAAG,GGGAAU,GGGAGG,GGGCGU,GGGCUG,GGGGAA,GGGGAG,GGGGCU,GGGGGC,GGGGGG,GGGUCG,GGGUGU,GGUCGA,GUCGAG,UCGAGG,UCGGGC,UGGGGG,UGGGGU,UGGGUG,UGUGGG,UUGGGU |
| SRSF10 |
6 |
13860 |
0.16666667 |
0.0200874160 |
3.052602 |
AAGAAG,AAGACA,AAGAGG,AGAGGA,GAGGAA |
AAAAGA,AAAGAA,AAAGAC,AAAGAG,AAAGGA,AAAGGG,AAGAAA,AAGAAG,AAGACA,AAGAGA,AAGAGG,AAGGAA,AAGGAG,AAGGGA,AAGGGG,ACAAAG,AGACAA,AGAGAA,AGAGAC,AGAGAG,AGAGGA,AGAGGG,CAAAGA,GAAAGA,GACAAA,GAGAAA,GAGAAC,GAGAAG,GAGACA,GAGACC,GAGACG,GAGAGA,GAGAGC,GAGAGG,GAGGAA,GAGGAG,GAGGGA,GAGGGG |
| SRSF6 |
10 |
21880 |
0.26190476 |
0.0317100317 |
3.046031 |
AAGAAG,AGAAGA,AGGAAG,GAAGAA,GAGGAA,GGAAGA |
AACCUG,AAGAAG,ACCGGG,ACCGUC,ACCUGG,AGAAGA,AGCACC,AGCGGA,AGGAAG,AUCAAC,AUCCUG,AUCGUA,CAACCU,CACACG,CACAGG,CACCUG,CACGGA,CACUCG,CACUGG,CAGCAC,CAUCCU,CCACAC,CCACAG,CCACUC,CCACUG,CCCGGC,CCUCAC,CCUCAG,CCUCUC,CCUCUG,CCUGGC,CGCGUC,CUACAC,CUACAG,CUACUC,CUACUG,CUCACG,CUCAGG,CUCAUC,CUCUCG,CUCUGG,CUUCAC,CUUCAG,CUUCUC,CUUCUG,GAAGAA,GACGUC,GAGGAA,GAUCAA,GCACCU,GCAGCA,GCCGGA,GCCGUC,GCUCAU,GGAAGA,UACACG,UACAGG,UACGUC,UACUCG,UACUGG,UCAACC,UCACAC,UCACAG,UCACUC,UCACUG,UCAUCC,UCCGGA,UCCUGG,UCUCAC,UCUCAG,UCUCUC,UCUCUG,UGCGGA,UGCGGC,UGCGUA,UGCGUC,UGCGUG,UGUGGA,UUACAC,UUACAG,UUACUC,UUACUG,UUCACG,UUCAGG,UUCUCG,UUCUGG,UUUCAC,UUUCAG,UUUCUC,UUUCUG |
| NOVA2 |
1 |
4013 |
0.04761905 |
0.0058171047 |
3.033166 |
GAGUCA |
AACACC,ACCACC,AGACAU,AGAUCA,AGCACC,AGGCAU,AGUCAU,AUCAAC,AUCACC,AUCAUC,CCUAGA,CUAGAU,GAGACA,GAGGCA,GAGUCA,GAUCAC,GGGGGG,UAGAUC,UUUUUU |
| SFPQ |
10 |
24050 |
0.26190476 |
0.0348548043 |
2.909613 |
AAGAGG,AGAGGA,GAAGAA,GAAGAG,GAGGAA,GGAAGA,GGAGGA |
AAGAAC,AAGAGC,AAGAGG,AAGCAA,AAGGAA,AAGGAC,AAGGGA,ACUGGG,AGAGAG,AGAGGA,AGAGGU,AGAUCG,AGGAAC,AGGACC,AGGGAU,AGGGGG,AUCGGA,CAGGCA,CUAAGG,CUGGAG,CUGGGA,GAAGAA,GAAGAG,GAAGCA,GAAGGA,GACUGG,GAGAGG,GAGGAA,GAGGAC,GAGGUA,GAUCGG,GCAGGC,GGAAGA,GGACUG,GGAGAG,GGAGGA,GGAGGG,GGGAUC,GGGGAC,GGGGAU,GGGGGA,GGGGGG,GGUAAG,GGUCUG,GUAAGA,GUAAUG,GUAAUU,GUAGUG,GUAGUU,GUCUGG,GUGAUG,GUGAUU,GUGGUG,GUGGUU,UAAGAG,UAAGGA,UAAGGG,UAAUGG,UAAUGU,UAAUUG,UAAUUU,UAGAGA,UAGAUC,UAGGGG,UAGUGG,UAGUGU,UAGUUG,UAGUUU,UCGGAA,UCUAAG,UCUGGA,UGAAGC,UGAUGG,UGAUGU,UGAUUG,UGAUUU,UGCAGG,UGGAAG,UGGAGA,UGGAGC,UGGAGG,UGGUCU,UGGUGG,UGGUGU,UGGUUG,UGGUUU,UUAAUG,UUAAUU,UUAGUG,UUAGUU,UUGAAG,UUGAUG,UUGAUU,UUGGUG,UUGGUU,UUUUUU |
| HNRNPA1 |
7 |
18070 |
0.19047619 |
0.0261885646 |
2.862602 |
AAGAGG,AGAGGA,AGGAAG,GAGGAA,GGAGGA |
AAAAAA,AAAGAG,AAGAGG,AAGGAG,AAGGUG,AAGUAC,AAUUUA,ACUAGA,AGACUA,AGAGGA,AGAUAU,AGAUUA,AGGAAG,AGGAGC,AGGGAC,AGGGCA,AGGGUU,AGUACG,AGUAGG,AGUGAA,AGUGGU,AGUUAG,AUAGAA,AUAGCA,AUAGGG,AUUAGA,AUUUAA,AUUUAU,CAAAGA,CAAGGA,CAGGGA,CCAAGG,CCCCCC,CGUAGG,CUAGAC,GAAGGU,GACUAG,GACUUA,GAGGAA,GAGGAG,GAGUGG,GAUUAG,GCAGGC,GCCAAG,GGAAGG,GGACUU,GGAGCC,GGAGGA,GGCAGG,GGGACU,GGGCAG,GGGUUA,GGUGCG,GGUUAG,GUAAGU,GUACGC,GUAGGG,GUAGGU,GUGGUG,GUUAGG,GUUUAG,UAAGUA,UACGGC,UAGACA,UAGACU,UAGAGA,UAGAGU,UAGAUU,UAGGAA,UAGGAU,UAGGCA,UAGGCU,UAGGGA,UAGGGC,UAGGGU,UAGGUA,UAGGUC,UAGUGA,UAGUUA,UCGGGC,UGGUGC,UUAGAU,UUAGGG,UUAUUU,UUUAGA,UUUAUA,UUUAUU,UUUUUU |
| FMR1 |
5 |
20291 |
0.14285714 |
0.0294072466 |
2.280330 |
AGGAAG,AGGAUG,GAGGAU,UGAGGA |
AAAAAA,AAGCGG,AAGGAA,AAGGAG,AAGGAU,AAGGGA,ACUAAG,ACUGGG,ACUGGU,AGCAGC,AGCGAA,AGCGAC,AGCGGC,AGCUGG,AGGAAG,AGGAAU,AGGAGG,AGGAGU,AGGAUG,AGGAUU,AGGCAC,AGGGAG,AGUGGC,AUAGGC,AUGGAG,CAGCUG,CAUAGG,CGAAGG,CGACUG,CGGCUG,CUAAGG,CUAUGG,CUGAGG,CUGGGC,CUGGUG,GAAGGA,GAAGGG,GACAAG,GACAGG,GACUAA,GACUGG,GAGCGA,GAGCGG,GAGGAU,GCAGCU,GCAUAG,GCGAAG,GCGACU,GCGGCU,GCUAAG,GCUAUG,GCUGAG,GCUGGG,GGACAA,GGACAG,GGACUA,GGCAUA,GGCGAA,GGCUAA,GGCUAU,GGCUGA,GGCUGG,GGGCAU,GGGCGA,GGGCUA,GGGCUG,GGGGGG,GUGCGA,GUGCGG,GUGGCU,UAAGGA,UAGCAG,UAGCGA,UAGCGG,UAGGCA,UAGUGG,UAUGGA,UGACAA,UGACAG,UGAGGA,UGCGAC,UGCGGC,UGGAGU,UGGCUG,UUUUUU |
| HNRNPH3 |
1 |
7688 |
0.04761905 |
0.0111429292 |
2.095410 |
GGAGGA |
AAGGGA,AAGGUG,AAUGUG,AGGGAA,AGGGGA,AUCGGG,AUGUGG,AUUGGG,CGAGGG,CGGGCG,CGGGGG,GAAGGG,GAAUGU,GAGGGG,GGAAGG,GGAAUG,GGAGGA,GGAGGG,GGGAAG,GGGAAU,GGGAGG,GGGCGU,GGGCUG,GGGGAG,GGGGCU,GGGGGC,GGGGGG,GGGUCG,GGGUGU,GGUCGA,GUCGAG,UCGAGG,UCGGGC,UGGGUG,UGUGGG,UUGGGU |
| HNRNPA2B1 |
1 |
8607 |
0.04761905 |
0.0124747476 |
1.932528 |
AAGAAG |
AAGAAG,AAGGAA,AAGGAG,AAGGGG,AAUUUA,ACUAGA,AGAAGC,AGACUA,AGAUAU,AGGAAC,AGGAGC,AGGGGC,AGUAGG,AUAGCA,AUAGGG,AUUUAA,CAAGAA,CAAGGA,CAAGGG,CCAAGA,CCAAGG,CGAAGG,CUAGAC,GAAGCC,GAAGGA,GACUAG,GCCAAG,GCGAAG,GGAACC,GGAGCC,GGGGCC,GUAGGG,UAGACA,UAGACU,UAGGGA,UAGGGU,UUAGGG,UUUAUA |
| SRSF1 |
22 |
99563 |
0.54761905 |
0.1442885423 |
1.924216 |
AAGAAG,AAGAGG,ACAGUG,AGAAGA,AGACAG,AGAGGA,AGGAAG,CGGAGG,GAAGAA,GAAGAC,GAAGAG,GAGGAA,GAGGAU,GCGGAG,GGAAGA,GGAGGA,GGAUGA,GGCGGA |
AAAAGA,AAAGAA,AAAGAC,AAAGAG,AAAGGA,AACACG,AACAGC,AACCGG,AAGAAC,AAGAAG,AAGACC,AAGAGA,AAGAGC,AAGAGG,AAGGAC,AAGGAG,AAGGCG,AAGGUG,AAUGAC,AAUUUC,ACAAGG,ACACGA,ACACGU,ACAGAA,ACAGAG,ACAGCA,ACAGCG,ACAGGA,ACAGGG,ACAGGU,ACAGUG,ACCCGA,ACCCGG,ACCCGU,ACCGGA,ACCGGU,ACGAAC,ACGAAG,ACGAAU,ACGACG,ACGCGA,ACGCGC,ACGCUC,ACGGAA,ACGGAC,ACUGAG,ACUGGA,AGAAGA,AGAAGG,AGACAA,AGACAG,AGACGA,AGACGU,AGAGAA,AGAGAC,AGAGCA,AGAGGA,AGAGGG,AGAGGU,AGAUGG,AGCAGG,AGCCGA,AGCCGU,AGCGAG,AGCGGA,AGCGGG,AGCGGU,AGGAAA,AGGAAC,AGGAAG,AGGACA,AGGACC,AGGACG,AGGACU,AGGAGA,AGGAGC,AGGAGG,AGGCGA,AGGUAA,AGUCGG,AUACGA,AUGAAC,AUGAAG,AUGACA,AUGACU,AUGGAC,AUGGAG,AUUACG,AUUUCA,CAACCA,CAACCC,CAACCG,CAACGA,CAACGC,CAAGCA,CAAGGA,CAAUGG,CACACG,CACAGA,CACAGC,CACAGG,CACAGU,CACCCA,CACCCG,CACCGA,CACCGG,CACGCA,CACGCU,CACGGA,CACUGG,CAGAAC,CAGACA,CAGACG,CAGAGA,CAGAGC,CAGAGG,CAGCCG,CAGCGG,CAGUCG,CAUGGU,CCAACC,CCAACG,CCAAGC,CCAAGG,CCAAUG,CCACAG,CCACCA,CCACCC,CCACCG,CCACGA,CCACGC,CCACGG,CCACUG,CCAGCA,CCAGCC,CCAGCG,CCAGGA,CCAGGG,CCAUGA,CCAUGG,CCCACC,CCCACG,CCCACU,CCCAGC,CCCAGG,CCCAUG,CCCCCA,CCCCCC,CCCCCG,CCCCGA,CCCCGC,CCCCGG,CCCGAC,CCCGCA,CCCGCC,CCCGCG,CCCGGA,CCCGGC,CCCGGG,CCCGUU,CCCUCC,CCCUCG,CCCUGA,CCCUGC,CCCUGG,CCGACA,CCGACC,CCGACG,CCGACU,CCGAGC,CCGAGG,CCGAUG,CCGCCA,CCGCCC,CCGCCG,CCGCGA,CCGCGC,CCGCGG,CCGCUA,CCGGAC,CCGGAG,CCGGCA,CCGGCC,CCGGCG,CCGGGA,CCGGGC,CCGGGG,CCGUCC,CCGUCG,CCGUGC,CCGUGG,CCGUUU,CCUAGG,CCUCCA,CCUCCG,CCUCGA,CCUCGG,CCUGAC,CCUGCA,CCUGCG,CCUGGA,CCUGGG,CGAACC,CGAACG,CGAAGC,CGAAGG,CGACAG,CGACCA,CGACCC,CGACCG,CGACGA,CGACGC,CGACGG,CGAGCA,CGAGCG,CGAGGA,CGAGGC,CGAGGG,CGAUGA,CGAUGG,CGCACA,CGCACC,CGCACG,CGCAGC,CGCAGG,CGCCCA,CGCCCC,CGCCCG,CGCCGA,CGCCGC,CGCCGG,CGCCGU,CGCGAG,CGCGCA,CGCGCC,CGCGCG,CGCGCU,CGCGGA,CGCGGC,CGCGGG,CGCGGU,CGCGUC,CGCUAU,CGCUCA,CGCUCC,CGCUCG,CGCUGC,CGCUGG,CGGAAC,CGGAAG,CGGAAU,CGGACA,CGGACC,CGGACG,CGGAGC,CGGAGG,CGGCAC,CGGCAG,CGGCCA,CGGCCC,CGGCCG,CGGCGA,CGGCGC,CGGCGG,CGGGCA,CGGGCC,CGGGCG,CGGGGA,CGGGGC,CGGGGG,CGGUCC,CGGUCG,CGGUGC,CGGUGG,CGUCCA,CGUCCG,CGUCGA,CGUCGG,CGUGCA,CGUGCG,CGUGGA,CGUGGG,CUACGA,CUAGGG,CUCAGG,CUCCGG,CUCGGA,CUCGUG,CUGAAC,CUGACU,CUGAGC,CUGAGU,GAAAGA,GAAAGG,GAACAG,GAACCA,GAACGA,GAAGAA,GAAGAC,GAAGAG,GAAGAU,GAAGCA,GAAGCC,GAAGCU,GAAGGA,GAAGGC,GAAGGU,GAAUGA,GACAGA,GACAGG,GACCCA,GACCCG,GACCGA,GACGAA,GACGAC,GACGAG,GACGCA,GACGCG,GACGGA,GACGGC,GACGGG,GACUGA,GACUGG,GAGAAC,GAGAAG,GAGACA,GAGACG,GAGCAG,GAGCGA,GAGCGG,GAGGAA,GAGGAC,GAGGAG,GAGGAU,GAGGCA,GAGGGA,GAGGGC,GAGGGG,GAGGUA,GAUACG,GAUGAA,GAUGAC,GAUGAU,GAUGCA,GAUGCC,GAUGCU,GAUGGA,GAUGGC,GAUGGU,GCACCA,GCACCG,GCACGA,GCACGG,GCACGU,GCAGCA,GCAGCG,GCAGGA,GCAGGC,GCAGGG,GCAGGU,GCAUAC,GCCCAC,GCCCCA,GCCCCG,GCCCGA,GCCCGG,GCCCGU,GCCGCA,GCCGCG,GCCGGA,GCCGGG,GCCGGU,GCCGUA,GCGAGC,GCGCAA,GCGCCA,GCGCCC,GCGCCG,GCGCGA,GCGCGC,GCGCGG,GCGGAA,GCGGAC,GCGGAG,GCGGCA,GCGGCG,GCGGGA,GCGGGG,GCGGUU,GCGUCA,GCUCCA,GCUCCG,GCUCGA,GCUCGG,GCUGCA,GCUGCG,GCUGGA,GCUGGG,GGAAAG,GGAACA,GGAAGA,GGAAGG,GGAAUG,GGACAA,GGACAG,GGACCA,GGACCG,GGACGA,GGACGG,GGACGU,GGACUG,GGAGAA,GGAGAC,GGAGAU,GGAGCA,GGAGCC,GGAGCG,GGAGCU,GGAGGA,GGAGGC,GGAGGG,GGAGGU,GGAUAC,GGAUAU,GGAUGA,GGAUUC,GGCACA,GGCACG,GGCAGA,GGCCCA,GGCCCG,GGCCGA,GGCCGG,GGCCGU,GGCGCA,GGCGCG,GGCGGA,GGCGGG,GGCGGU,GGGACG,GGGCCA,GGGCCG,GGGCGA,GGGCGG,GGGGAA,GGGGAC,GGGGAG,GGGGCA,GGGGCG,GGGGGA,GGGGGG,GGGUAC,GGUAAC,GGUCCA,GGUCCG,GGUCGA,GGUCGG,GGUGAA,GGUGAC,GGUGAU,GGUGCA,GGUGCC,GGUGCG,GGUGCU,GGUGGA,GGUGGC,GGUGGG,GGUGGU,GUAGGA,GUGACA,GUGACG,UAAUUU,UACGAA,UACGGA,UAGACA,UAGGAC,UAUUAC,UCAAGA,UCAGGU,UCCGGA,UCGCAC,UCGGGC,UCGUGU,UGAACA,UGAACC,UGAAGA,UGAAGC,UGAAGG,UGACAG,UGACGA,UGACUA,UGACUG,UGAGUU,UGAUGA,UGAUGC,UGAUGG,UGGAAA,UGGACA,UGGAGA,UGGAGC,UGGAGG,UGGUGA,UGGUGC,UGGUGG,UGUAGG,UUAAUU,UUACGG,UUCAAG,UUGACA,UUUCAA |
| HNRNPH2 |
1 |
9711 |
0.04761905 |
0.0140746688 |
1.758438 |
GGAGGA |
AAGGCG,AAGGGA,AAGGGG,AAGGUG,AAUGUG,AGGCGA,AGGGAA,AGGGGA,AUCGGG,AUGUGG,AUUGGG,CAGGAC,CGAGGG,CGGGCG,CGGGGG,CUGGGG,GAAGGG,GAAUGU,GAGGGG,GGAAGG,GGAAUG,GGAGGA,GGAGGG,GGCGAA,GGGAAG,GGGAAU,GGGAGG,GGGCGU,GGGCUG,GGGGAA,GGGGAG,GGGGCU,GGGGGA,GGGGGC,GGGGGG,GGGUCG,GGGUGU,GGUCGA,GUCGAG,UCGAGG,UCGGGC,UGGGGG,UGGGGU,UGGGUG,UGUGGG,UUGGGU |
| SRSF2 |
10 |
88240 |
0.26190476 |
0.1278792060 |
1.034261 |
AAGAAG,AGAAGA,AGAGGA,AGGAAG,GAAGAA,GAGGAA,GCGGAG,GGAGGA |
AAAAGA,AAAGAG,AAAGGA,AACCAG,AACGAG,AAGAAG,AAGAGA,AAGCAG,AAGCGG,AAGCUG,AAGGAG,AAGGCG,AAGUAC,AAUACC,AAUCCC,AAUGCC,AAUGCU,ACCACC,ACCACU,ACCAGC,ACCAGU,ACCCCC,ACCCCU,ACCCGC,ACCCGG,ACCGGU,ACCGUA,ACCUCC,ACCUGC,ACGAGA,ACGAGG,ACGCUG,ACGGAG,ACGUGU,ACUACC,ACUACU,ACUAGA,ACUCAA,ACUCCC,ACUCCU,ACUGUA,AGAAGA,AGAAGC,AGAAUA,AGAAUG,AGACUA,AGAGAA,AGAGAU,AGAGGA,AGAGGU,AGAGUA,AGAGUG,AGAGUU,AGAUAA,AGAUCC,AGAUGC,AGCACU,AGCAGA,AGCAGU,AGCCCC,AGCCGA,AGCCUC,AGCGGA,AGCGUG,AGCUCC,AGCUGU,AGGAAG,AGGAGA,AGGAGC,AGGAGG,AGGAGU,AGGCAG,AGGCGA,AGGCGU,AGGCUG,AGGGUA,AGUACG,AGUAGA,AGUAGG,AGUAGU,AGUAUC,AGUCGA,AGUGAC,AGUGGU,AGUGUU,AGUUCC,AUACCG,AUCACC,AUCACU,AUCCCC,AUCCCG,AUCCCU,AUCCGG,AUCGAG,AUCGCU,AUGAAU,AUGAUG,AUGCCG,AUGCGU,AUGCUG,AUGGAG,AUUACC,AUUACU,AUUAGU,AUUCCA,AUUCCC,AUUCCU,AUUGAU,CACCUC,CAGAGA,CAGAGG,CAGAGU,CAGCUA,CAGUAG,CAGUCC,CAGUGG,CAGUGU,CCACCA,CCACCG,CCACUA,CCACUG,CCAGAA,CCAGAU,CCAGCC,CCAGCU,CCAGGG,CCAGGU,CCAGUC,CCAGUG,CCAGUU,CCCACA,CCCCCA,CCCCCG,CCCCUA,CCCCUG,CCCGCA,CCCGCU,CCCGUG,CCCUUG,CCGAGA,CCGCAG,CCGCUA,CCGGAU,CCGGUG,CCUCAG,CCUCCG,CCUGCG,CCUGCU,CCUGUU,CGAACG,CGAGAA,CGAGAG,CGAGAU,CGAGCA,CGAGGA,CGAGUA,CGAGUG,CGAGUU,CGCAGG,CGCAGU,CGCCCU,CGCUGC,CGGAGA,CGGAGU,CGUAAG,CGUGAU,CGUGCG,CGUGUA,CGUGUU,CUAACG,CUACCA,CUACCG,CUACUA,CUACUG,CUAGAA,CUAGAC,CUCAAG,CUCCAA,CUCCCA,CUCCCG,CUCCUA,CUCCUG,CUCGAG,CUCGUG,CUGAUG,CUGCAA,CUGCUA,CUGUUA,GAAAGG,GAACCA,GAACGA,GAAGAA,GAAGCG,GAAGGC,GAAUAC,GAAUCC,GAAUGC,GACCAC,GACCCC,GACCCG,GACCGG,GACCGU,GACCUG,GACGCU,GACGGA,GACUAC,GACUAG,GACUCA,GACUCC,GACUGU,GAGAAG,GAGAAU,GAGAGA,GAGAGG,GAGAGU,GAGAUA,GAGAUG,GAGCAC,GAGCAG,GAGCGU,GAGCUG,GAGGAA,GAGGAC,GAGGAG,GAGUAU,GAGUGA,GAUCAC,GAUCCC,GAUCCG,GAUCGA,GAUCGC,GAUGAU,GAUGCG,GAUGCU,GAUGGA,GAUUAC,GAUUAG,GAUUCC,GAUUGA,GCACUG,GCAGAG,GCAGGG,GCAGGU,GCAGUA,GCAGUG,GCCAAA,GCCACC,GCCACU,GCCCAC,GCCCCC,GCCCCU,GCCCUU,GCCGAG,GCCGCA,GCCGCC,GCCGUU,GCCUCA,GCCUCC,GCGAGC,GCGAGU,GCGCAG,GCGGAG,GCGGUC,GCGUGU,GCUACC,GCUACU,GCUCCA,GCUCCC,GCUCCU,GCUCGA,GCUCGU,GCUGAU,GCUGCA,GCUGCU,GCUGUU,GGAACC,GGAAGG,GGAAUG,GGACCG,GGACGC,GGACUG,GGAGAA,GGAGAG,GGAGAU,GGAGCC,GGAGCG,GGAGGA,GGAGUA,GGAGUG,GGAGUU,GGAUCC,GGAUGG,GGCAGU,GGCCAA,GGCCAC,GGCCCA,GGCCCC,GGCCGC,GGCCUC,GGCGAG,GGCGUG,GGCUAC,GGCUCC,GGCUCG,GGCUGA,GGCUGC,GGGAAU,GGGAGC,GGGCAG,GGGUAA,GGGUAC,GGGUAG,GGGUAU,GGGUGA,GGUAAG,GGUACG,GGUAGG,GGUAUG,GGUCAC,GGUCAG,GGUCCC,GGUCGC,GGUGAG,GGUGAU,GGUUAA,GGUUAC,GGUUCC,GGUUGG,GGUUGU,GUAACC,GUAAGC,GUAAGU,GUACGA,GUACGC,GUAGGC,GUAGGG,GUAGUC,GUAUCC,GUAUGC,GUCACC,GUCACU,GUCAGU,GUCCCC,GUCCCU,GUCCUC,GUCGAU,GUCGCC,GUCUAA,GUGAGC,GUGAUG,GUGCAG,GUGGUC,GUGUAA,GUUAAU,GUUACC,GUUACG,GUUACU,GUUCCA,GUUCCC,GUUCCG,GUUCCU,GUUCGA,GUUCGU,GUUCUG,GUUGGC,GUUGUU,GUUUCG,UAACCA,UAAGCU,UAAGUA,UACCAG,UACGAG,UACGCU,UACGUG,UAGACU,UAGGCU,UAUCCC,UAUGCC,UAUGCU,UCACCA,UCACCG,UCACUA,UCACUG,UCAGUG,UCCAGA,UCCAGC,UCCAGG,UCCAGU,UCCCCA,UCCCCG,UCCCCU,UCCCGC,UCCCGU,UCCCUA,UCCCUG,UCCGGA,UCCUCA,UCCUCC,UCCUGC,UCCUGU,UCGAGA,UCGAGU,UCGCCG,UCGCUG,UCGUAA,UCGUGC,UCUAAC,UCUGUA,UGAAUG,UGAGCU,UGAUCG,UGAUGC,UGAUGG,UGCAGA,UGCAGU,UGCCGC,UGCCGU,UGCGGU,UGCUGU,UGGAGA,UGGAGG,UGGAGU,UGGCAG,UGUUAC,UGUUCC,UGUUCG,UUAAUG,UUACCA,UUACCG,UUACGU,UUACUA,UUACUG,UUAGUG,UUCCAG,UUCCCA,UUCCCG,UUCCCU,UUCCGG,UUCCUA,UUCCUG,UUCGAG,UUCGUG,UUCUGU,UUGAUG,UUGGCG,UUGUUG,UUUCGA |