circRNA Analysis
WANG Ziyi
Date: 18 5月 2022
ENSG00000170144:+:2:177217704:177217845
1. Sequences
| id | gene | transcript | strand | chrom | startGR | endGR | length | seq | type |
|---|---|---|---|---|---|---|---|---|---|
| ENSG00000170144:+:2:177217704:177217845 | ENSG00000170144 | ENST00000676762 | + | 2 | 177217705 | 177217845 | 141 | GUGGCAACUAUGGCGGUGGUCCUGGUUAUAGUAGUAGAGGGGGCUAUGGUGGUGGUGGACCAGGAUAUGGAAACCAAGGUGGUGGAUAUGGUGGAGGUGGAGGAUAUGAUGGUUACAAUGAAGGAGGAAAUUUUGGCGGUGGUGGCAACUAUGGCGGUGGUCCUGGUUAUAGUAGUAGAGGGGGCUAUGGU | circ |
| ENSG00000170144:+:2:177217704:177217845 | ENSG00000170144 | ENST00000676762 | + | 2 | 177217705 | 177217845 | 22 | UUUUGGCGGUGGUGGCAACUAU | bsj |
| ENSG00000170144:+:2:177217704:177217845 | ENSG00000170144 | ENST00000676762 | + | 2 | 177217505 | 177217714 | 210 | CCAUAGUCUAGCUACUCAGGAGACUGAGGUGGGAGAAUGGCUUGAGCCCAGGAGUUCAAGGCUGCAUUGCUGUGGUUGCACCACUGUACUUGAGCCCGGGCAACAGAGCAAGACCCAGUCUCUUACACACACACCAGAAUUGAAUAACGUGACUAUACACUUAAGUUUUUGUGUGGUCUUGUUAUUUGUUGUUUUUUUAGGUGGCAACUA | ie_up |
| ENSG00000170144:+:2:177217704:177217845 | ENSG00000170144 | ENST00000676762 | + | 2 | 177217836 | 177218045 | 210 | UUUGGCGGUGGUAAGCAUUCACUUGUUUUAUUUAAAUGUUAAAUAUUCAGUGUUGCUAACAGUUCCCAUGACACAUAUUUGGAAAGUGUUAAAAGCGUUUGAUCAAAUUUUAUGUUCUAUAAGAAAAAUACUAAAAUGGUUGGUAGUUCAAACCAAAUUUUCUUGACUUUGCUGGUUAUUUAGUAAAAUGUACAAAUGUACUCAGCUCUC | ie_down |
- Note:
- id: unique identifier.
- gene: represents the gene from which the circRNA arises.
- transcript: transcript whose exon coordinates overlap with the detected back-spliced junction coordinate and used in the downstream analysis.
- strand: is the strand from which the gene is transcribed.
- chrom: is the chromosome from which the circRNA is derived.
- startGR: is the 5’ coordinate of the genomic range from which the sequence are extracted.
- endGR: is the 3’ coordinate of the genomic range from which the sequence are extracted.
- length: length of the extracted sequences.
- seq: sequence used in the downstream analysis.
- type: type of sequences retrieved. If type = “circ” the sequences derive from the internal circRNA sequences. If type = “bsj” the sequences derive from the back spliced junctions (BSJ). If type = “ie_up” the Intron or Exon sequences derive from the up-stream of BSJ up to 210 bp. If type = “ie_down” the Intron or Exon sequences derive from the down-stream of BSJ up to 210 bp.
2. RNA Binding Proteins Analysis
RBP on full sequence
Plot
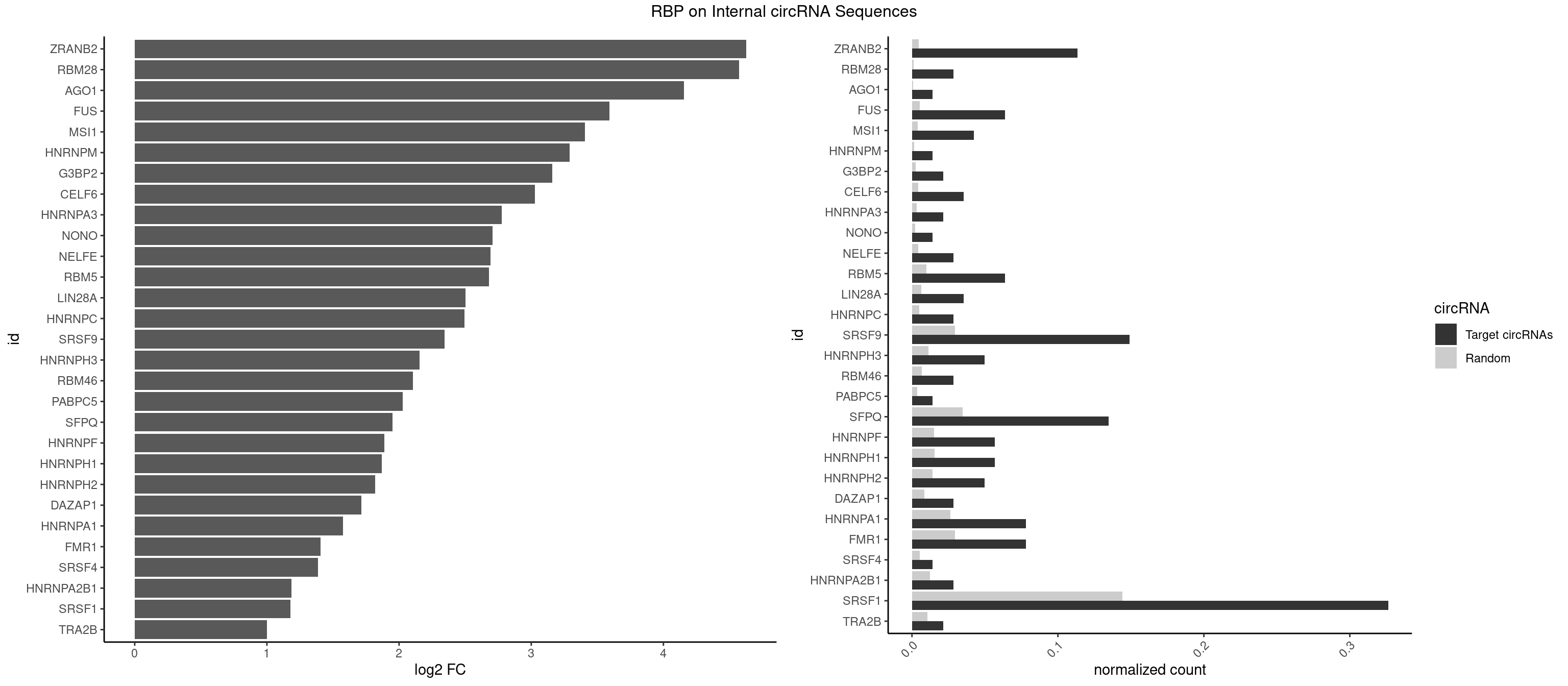
Spreadsheet
| id | foreground | background | foregroundNorm | backgroundNorm | log2FC | motifF | motifB |
|---|---|---|---|---|---|---|---|
| ZRANB2 | 15 | 3173 | 0.11347518 | 0.004599773 | 4.624670 | GGUGGU,GUGGUG,UGGUGG | AAAGGU,AGGGAA,AGGUAA,AGGUAC,AGGUAG,AGGUAU,AGGUCU,AGGUUA,AGGUUU,CGGUAA,CGGUAG,CGGUAU,CGGUCU,GAGGUU,GGGUAA,GGGUGA,GGUAAA,GGUGGU,GUAAAG,GUGGUG,UAAAGG,UGGUAA,UGGUGG |
| RBM28 | 3 | 822 | 0.02836879 | 0.001192695 | 4.572008 | AGUAGA,AGUAGU | AGUAGA,AGUAGG,AGUAGU,GAGUAG,GUGUAG,UGUAGA,UGUAGG,UGUAGU |
| AGO1 | 1 | 548 | 0.01418440 | 0.000795613 | 4.156094 | GUAGUA | AGGUAG,GAGGUA,GGUAGU,GUAGUA,UGAGGU |
| FUS | 8 | 3648 | 0.06382979 | 0.005288145 | 3.593396 | CGGUGG,UGGUGG | AAAAAA,CGGUGA,CGGUGG,GGGGGG,GGGUGA,GGGUGC,GGGUGG,GGGUGU,UGGUGA,UGGUGG,UGGUGU,UUUUUU |
| MSI1 | 5 | 2770 | 0.04255319 | 0.004015744 | 3.405528 | AGGAGG,AGGUGG,UAGUAG | AGGAAG,AGGAGG,AGGUAG,AGGUGG,AGUAAG,AGUAGG,AGUUAG,AGUUGG,UAGGAA,UAGGAG,UAGGUA,UAGGUG,UAGUAA,UAGUAG,UAGUUA,UAGUUG |
| HNRNPM | 1 | 999 | 0.01418440 | 0.001449204 | 3.290972 | GAAGGA | AAGGAA,GAAGGA,GGGGGG |
| G3BP2 | 2 | 1644 | 0.02127660 | 0.002383941 | 3.157847 | AGGAUA | AGGAUA,AGGAUG,AGGAUU,GGAUAA,GGAUAG,GGAUGA,GGAUGG,GGAUUA,GGAUUG |
| CELF6 | 4 | 2997 | 0.03546099 | 0.004344713 | 3.028900 | GUGGUG | GUGAGG,GUGAUG,GUGGGG,GUGGUG,GUGUGG,GUGUUG,UGUGAG,UGUGAU,UGUGGG,UGUGGU,UGUGUG,UGUGUU |
| HNRNPA3 | 2 | 2140 | 0.02127660 | 0.003102746 | 2.777650 | AAGGAG,CCAAGG | AAGGAG,AGGAGC,CAAGGA,CCAAGG,GCCAAG,GGAGCC |
| NONO | 1 | 1498 | 0.01418440 | 0.002172357 | 2.706972 | GAGGAA | AGAGGA,AGGAAC,GAGAGG,GAGGAA |
| NELFE | 3 | 3028 | 0.02836879 | 0.004389639 | 2.692131 | CUGGUU,UGGUUA | CUCUCU,CUCUGG,CUGGCU,CUGGUU,GCUAAC,GGCUAA,GGUCUC,GGUUAG,GUCUCU,UCUCUC,UCUCUG,UCUGGC,UCUGGU,UGGCUA,UGGUUA |
| RBM5 | 8 | 6879 | 0.06382979 | 0.009970523 | 2.678489 | AAGGAG,AAGGUG,GAAGGA,GGUGGU | AAAAAA,AAGGAA,AAGGAG,AAGGGG,AAGGUA,AAGGUG,AGGGAA,AGGGAG,AGGGUA,AGGGUG,AGGUAA,CAAGGA,CAAGGG,CCCCCC,CUCUUC,CUUCUC,GAAGGA,GAAGGG,GAAGGU,GAGGGA,GAGGGU,GGGGGG,GGUGGU,UCUUCU,UUCUCU |
| LIN28A | 4 | 4315 | 0.03546099 | 0.006254764 | 2.503206 | GGAGGA,UGGAGG | AGGAGA,AGGAGU,CGGAGA,CGGAGG,CGGAGU,GGAGAA,GGAGAU,GGAGGA,GGAGGG,GGAGUA,UGGAGA,UGGAGG,UGGAGU |
| HNRNPC | 3 | 3471 | 0.02836879 | 0.005031636 | 2.495205 | GGAUAU | AUUUUU,CUUUUU,GCAUAC,GGAUAC,GGAUAU,GGAUUC,GGGGGG,GGGUAC,UUUUUA,UUUUUC,UUUUUG,UUUUUU |
| SRSF9 | 20 | 20254 | 0.14893617 | 0.029353626 | 2.343084 | AGGAAA,GAAGGA,GGACCA,GGAGGA,GGUGGA,GGUGGC,UGAAGG,UGAUGG,UGGAGG,UGGUGG | AGGAAA,AGGAAC,AGGACA,AGGACC,AGGAGA,AGGAGC,AUGAAA,AUGAAC,AUGACA,AUGACC,AUGAGA,AUGAGC,CUGGAU,GAAGCA,GAAGCC,GAAGGA,GAAGGC,GAUGCA,GAUGCC,GAUGGA,GAUGGC,GGAAAA,GGAAAG,GGAACA,GGAACG,GGAAGC,GGAAGG,GGACAA,GGACAG,GGACCA,GGACCG,GGAGAA,GGAGAG,GGAGCA,GGAGCC,GGAGCG,GGAGGA,GGAGGC,GGAUGC,GGAUGG,GGGAGC,GGGAGG,GGGUGC,GGGUGG,GGUGCA,GGUGCC,GGUGGA,GGUGGC,UGAAAA,UGAAAG,UGAACA,UGAACG,UGAAGC,UGAAGG,UGACAA,UGACAG,UGACCA,UGACCG,UGAGAA,UGAGAG,UGAGCA,UGAGCG,UGAUGC,UGAUGG,UGGAGC,UGGAGG,UGGAUU,UGGUGC,UGGUGG |
| HNRNPH3 | 6 | 7688 | 0.04964539 | 0.011142929 | 2.155531 | AAGGUG,GAGGGG,GGAGGA,GGGGCU,GGGGGC | AAGGGA,AAGGUG,AAUGUG,AGGGAA,AGGGGA,AUCGGG,AUGUGG,AUUGGG,CGAGGG,CGGGCG,CGGGGG,GAAGGG,GAAUGU,GAGGGG,GGAAGG,GGAAUG,GGAGGA,GGAGGG,GGGAAG,GGGAAU,GGGAGG,GGGCGU,GGGCUG,GGGGAG,GGGGCU,GGGGGC,GGGGGG,GGGUCG,GGGUGU,GGUCGA,GUCGAG,UCGAGG,UCGGGC,UGGGUG,UGUGGG,UUGGGU |
| RBM46 | 3 | 4554 | 0.02836879 | 0.006601124 | 2.103521 | AAUGAA,AUGAAG,AUGAUG | AAUCAA,AAUCAU,AAUGAA,AAUGAU,AUCAAA,AUCAAG,AUCAAU,AUCAUA,AUCAUG,AUCAUU,AUGAAA,AUGAAG,AUGAAU,AUGAUA,AUGAUG,AUGAUU,GAUCAA,GAUCAU,GAUGAA,GAUGAU |
| PABPC5 | 1 | 2400 | 0.01418440 | 0.003479539 | 2.027337 | GAAAUU | AGAAAA,AGAAAG,AGAAAU,GAAAAU,GAAAGU,GAAAUU |
| SFPQ | 18 | 24050 | 0.13475177 | 0.034854804 | 1.950875 | AGGGGG,GAAGGA,GAGGAA,GGAGGA,GUGGUG,UGAUGG,UGGAGG,UGGUGG | AAGAAC,AAGAGC,AAGAGG,AAGCAA,AAGGAA,AAGGAC,AAGGGA,ACUGGG,AGAGAG,AGAGGA,AGAGGU,AGAUCG,AGGAAC,AGGACC,AGGGAU,AGGGGG,AUCGGA,CAGGCA,CUAAGG,CUGGAG,CUGGGA,GAAGAA,GAAGAG,GAAGCA,GAAGGA,GACUGG,GAGAGG,GAGGAA,GAGGAC,GAGGUA,GAUCGG,GCAGGC,GGAAGA,GGACUG,GGAGAG,GGAGGA,GGAGGG,GGGAUC,GGGGAC,GGGGAU,GGGGGA,GGGGGG,GGUAAG,GGUCUG,GUAAGA,GUAAUG,GUAAUU,GUAGUG,GUAGUU,GUCUGG,GUGAUG,GUGAUU,GUGGUG,GUGGUU,UAAGAG,UAAGGA,UAAGGG,UAAUGG,UAAUGU,UAAUUG,UAAUUU,UAGAGA,UAGAUC,UAGGGG,UAGUGG,UAGUGU,UAGUUG,UAGUUU,UCGGAA,UCUAAG,UCUGGA,UGAAGC,UGAUGG,UGAUGU,UGAUUG,UGAUUU,UGCAGG,UGGAAG,UGGAGA,UGGAGC,UGGAGG,UGGUCU,UGGUGG,UGGUGU,UGGUUG,UGGUUU,UUAAUG,UUAAUU,UUAGUG,UUAGUU,UUGAAG,UUGAUG,UUGAUU,UUGGUG,UUGGUU,UUUUUU |
| HNRNPF | 7 | 10561 | 0.05673759 | 0.015306492 | 1.890161 | AAGGUG,GAGGAA,GAGGGG,GGAGGA,GGGGCU,GGGGGC | AAGGCG,AAGGGA,AAGGGG,AAGGUG,AAUGUG,AGGAAG,AGGCGA,AGGGAA,AGGGAU,AGGGGA,AUGGGA,AUGGGG,AUGUGG,CGAUGG,CGGGAU,CGGGGG,CUGGGG,GAAGGG,GAAGGU,GAAUGU,GAGGAA,GAGGGG,GAUGGG,GGAAGG,GGAAUG,GGAGGA,GGAGGG,GGAUGG,GGCGAA,GGGAAG,GGGAAU,GGGAGG,GGGAUG,GGGCUG,GGGGAA,GGGGAG,GGGGCU,GGGGGC,GGGGGG,GGGGUA,UGGGAA,UGGGGU,UGUGGG |
| HNRNPH1 | 7 | 10717 | 0.05673759 | 0.015532568 | 1.869008 | AAGGUG,GAGGAA,GAGGGG,GGAGGA,GGGGCU,GGGGGC | AAGGCG,AAGGGA,AAGGGG,AAGGUG,AAUGUG,AGGAAG,AGGCGA,AGGGAA,AGGGGA,AUCGGG,AUGUGG,AUUGGG,CAGGAC,CGAGGG,CGGGCG,CGGGGG,CUGGGG,GAAGGG,GAAGGU,GAAUGU,GAGGAA,GAGGGG,GGAAGG,GGAAUG,GGAGGA,GGAGGG,GGCGAA,GGGAAG,GGGAAU,GGGAGG,GGGCGU,GGGCUG,GGGGAA,GGGGAG,GGGGCU,GGGGGC,GGGGGG,GGGUCG,GGGUGU,GGUCGA,GUCGAG,UCGAGG,UCGGGC,UGGGGG,UGGGGU,UGGGUG,UGUGGG,UUGGGU |
| HNRNPH2 | 6 | 9711 | 0.04964539 | 0.014074669 | 1.818559 | AAGGUG,GAGGGG,GGAGGA,GGGGCU,GGGGGC | AAGGCG,AAGGGA,AAGGGG,AAGGUG,AAUGUG,AGGCGA,AGGGAA,AGGGGA,AUCGGG,AUGUGG,AUUGGG,CAGGAC,CGAGGG,CGGGCG,CGGGGG,CUGGGG,GAAGGG,GAAUGU,GAGGGG,GGAAGG,GGAAUG,GGAGGA,GGAGGG,GGCGAA,GGGAAG,GGGAAU,GGGAGG,GGGCGU,GGGCUG,GGGGAA,GGGGAG,GGGGCU,GGGGGA,GGGGGC,GGGGGG,GGGUCG,GGGUGU,GGUCGA,GUCGAG,UCGAGG,UCGGGC,UGGGGG,UGGGGU,UGGGUG,UGUGGG,UUGGGU |
| DAZAP1 | 3 | 5964 | 0.02836879 | 0.008644502 | 1.714450 | AGGAAA,AGGAUA | AAAAAA,AAUUUA,AGAUAU,AGGAAA,AGGAAG,AGGAUA,AGGAUG,AGGUAA,AGGUAG,AGGUUA,AGGUUG,AGUAAA,AGUAAG,AGUAGG,AGUAUA,AGUAUG,AGUUAA,AGUUAG,AGUUUA,AGUUUG,GGGGGG,GUAACG,UAGGAA,UAGGAU,UAGGUA,UAGGUU,UAGUAA,UAGUAU,UAGUUA,UAGUUU,UUUUUU |
| HNRNPA1 | 10 | 18070 | 0.07801418 | 0.026188565 | 1.574799 | AAGGAG,AAGGUG,CCAAGG,GAGGAA,GGAGGA,GUGGUG | AAAAAA,AAAGAG,AAGAGG,AAGGAG,AAGGUG,AAGUAC,AAUUUA,ACUAGA,AGACUA,AGAGGA,AGAUAU,AGAUUA,AGGAAG,AGGAGC,AGGGAC,AGGGCA,AGGGUU,AGUACG,AGUAGG,AGUGAA,AGUGGU,AGUUAG,AUAGAA,AUAGCA,AUAGGG,AUUAGA,AUUUAA,AUUUAU,CAAAGA,CAAGGA,CAGGGA,CCAAGG,CCCCCC,CGUAGG,CUAGAC,GAAGGU,GACUAG,GACUUA,GAGGAA,GAGGAG,GAGUGG,GAUUAG,GCAGGC,GCCAAG,GGAAGG,GGACUU,GGAGCC,GGAGGA,GGCAGG,GGGACU,GGGCAG,GGGUUA,GGUGCG,GGUUAG,GUAAGU,GUACGC,GUAGGG,GUAGGU,GUGGUG,GUUAGG,GUUUAG,UAAGUA,UACGGC,UAGACA,UAGACU,UAGAGA,UAGAGU,UAGAUU,UAGGAA,UAGGAU,UAGGCA,UAGGCU,UAGGGA,UAGGGC,UAGGGU,UAGGUA,UAGGUC,UAGUGA,UAGUUA,UCGGGC,UGGUGC,UUAGAU,UUAGGG,UUAUUU,UUUAGA,UUUAUA,UUUAUU,UUUUUU |
| FMR1 | 10 | 20291 | 0.07801418 | 0.029407247 | 1.407565 | AAGGAG,AGGAGG,CUAUGG,GAAGGA,GAGGAU,GCUAUG,GGCUAU,GGGCUA,UAUGGA | AAAAAA,AAGCGG,AAGGAA,AAGGAG,AAGGAU,AAGGGA,ACUAAG,ACUGGG,ACUGGU,AGCAGC,AGCGAA,AGCGAC,AGCGGC,AGCUGG,AGGAAG,AGGAAU,AGGAGG,AGGAGU,AGGAUG,AGGAUU,AGGCAC,AGGGAG,AGUGGC,AUAGGC,AUGGAG,CAGCUG,CAUAGG,CGAAGG,CGACUG,CGGCUG,CUAAGG,CUAUGG,CUGAGG,CUGGGC,CUGGUG,GAAGGA,GAAGGG,GACAAG,GACAGG,GACUAA,GACUGG,GAGCGA,GAGCGG,GAGGAU,GCAGCU,GCAUAG,GCGAAG,GCGACU,GCGGCU,GCUAAG,GCUAUG,GCUGAG,GCUGGG,GGACAA,GGACAG,GGACUA,GGCAUA,GGCGAA,GGCUAA,GGCUAU,GGCUGA,GGCUGG,GGGCAU,GGGCGA,GGGCUA,GGGCUG,GGGGGG,GUGCGA,GUGCGG,GUGGCU,UAAGGA,UAGCAG,UAGCGA,UAGCGG,UAGGCA,UAGUGG,UAUGGA,UGACAA,UGACAG,UGAGGA,UGCGAC,UGCGGC,UGGAGU,UGGCUG,UUUUUU |
| SRSF4 | 1 | 3740 | 0.01418440 | 0.005421472 | 1.387548 | GAGGAA | AAGAAG,AGAAGA,AGGAAG,GAAGAA,GAGGAA,GGAAGA |
| HNRNPA2B1 | 3 | 8607 | 0.02836879 | 0.012474748 | 1.185294 | AAGGAG,CCAAGG,GAAGGA | AAGAAG,AAGGAA,AAGGAG,AAGGGG,AAUUUA,ACUAGA,AGAAGC,AGACUA,AGAUAU,AGGAAC,AGGAGC,AGGGGC,AGUAGG,AUAGCA,AUAGGG,AUUUAA,CAAGAA,CAAGGA,CAAGGG,CCAAGA,CCAAGG,CGAAGG,CUAGAC,GAAGCC,GAAGGA,GACUAG,GCCAAG,GCGAAG,GGAACC,GGAGCC,GGGGCC,GUAGGG,UAGACA,UAGACU,UAGGGA,UAGGGU,UUAGGG,UUUAUA |
| SRSF1 | 45 | 99563 | 0.32624113 | 0.144288542 | 1.176982 | AAGGAG,AAGGUG,AGAGGG,AGGAAA,AGGAGG,AUGAAG,CCAAGG,CCAGGA,CGGUGG,GAAGGA,GAGGAA,GAGGAU,GAGGGG,GAUGGU,GGACCA,GGAGGA,GGAGGU,GGAUAU,GGCGGU,GGUGGA,GGUGGC,GGUGGU,UGAAGG,UGAUGG,UGGAAA,UGGAGG,UGGUGG | AAAAGA,AAAGAA,AAAGAC,AAAGAG,AAAGGA,AACACG,AACAGC,AACCGG,AAGAAC,AAGAAG,AAGACC,AAGAGA,AAGAGC,AAGAGG,AAGGAC,AAGGAG,AAGGCG,AAGGUG,AAUGAC,AAUUUC,ACAAGG,ACACGA,ACACGU,ACAGAA,ACAGAG,ACAGCA,ACAGCG,ACAGGA,ACAGGG,ACAGGU,ACAGUG,ACCCGA,ACCCGG,ACCCGU,ACCGGA,ACCGGU,ACGAAC,ACGAAG,ACGAAU,ACGACG,ACGCGA,ACGCGC,ACGCUC,ACGGAA,ACGGAC,ACUGAG,ACUGGA,AGAAGA,AGAAGG,AGACAA,AGACAG,AGACGA,AGACGU,AGAGAA,AGAGAC,AGAGCA,AGAGGA,AGAGGG,AGAGGU,AGAUGG,AGCAGG,AGCCGA,AGCCGU,AGCGAG,AGCGGA,AGCGGG,AGCGGU,AGGAAA,AGGAAC,AGGAAG,AGGACA,AGGACC,AGGACG,AGGACU,AGGAGA,AGGAGC,AGGAGG,AGGCGA,AGGUAA,AGUCGG,AUACGA,AUGAAC,AUGAAG,AUGACA,AUGACU,AUGGAC,AUGGAG,AUUACG,AUUUCA,CAACCA,CAACCC,CAACCG,CAACGA,CAACGC,CAAGCA,CAAGGA,CAAUGG,CACACG,CACAGA,CACAGC,CACAGG,CACAGU,CACCCA,CACCCG,CACCGA,CACCGG,CACGCA,CACGCU,CACGGA,CACUGG,CAGAAC,CAGACA,CAGACG,CAGAGA,CAGAGC,CAGAGG,CAGCCG,CAGCGG,CAGUCG,CAUGGU,CCAACC,CCAACG,CCAAGC,CCAAGG,CCAAUG,CCACAG,CCACCA,CCACCC,CCACCG,CCACGA,CCACGC,CCACGG,CCACUG,CCAGCA,CCAGCC,CCAGCG,CCAGGA,CCAGGG,CCAUGA,CCAUGG,CCCACC,CCCACG,CCCACU,CCCAGC,CCCAGG,CCCAUG,CCCCCA,CCCCCC,CCCCCG,CCCCGA,CCCCGC,CCCCGG,CCCGAC,CCCGCA,CCCGCC,CCCGCG,CCCGGA,CCCGGC,CCCGGG,CCCGUU,CCCUCC,CCCUCG,CCCUGA,CCCUGC,CCCUGG,CCGACA,CCGACC,CCGACG,CCGACU,CCGAGC,CCGAGG,CCGAUG,CCGCCA,CCGCCC,CCGCCG,CCGCGA,CCGCGC,CCGCGG,CCGCUA,CCGGAC,CCGGAG,CCGGCA,CCGGCC,CCGGCG,CCGGGA,CCGGGC,CCGGGG,CCGUCC,CCGUCG,CCGUGC,CCGUGG,CCGUUU,CCUAGG,CCUCCA,CCUCCG,CCUCGA,CCUCGG,CCUGAC,CCUGCA,CCUGCG,CCUGGA,CCUGGG,CGAACC,CGAACG,CGAAGC,CGAAGG,CGACAG,CGACCA,CGACCC,CGACCG,CGACGA,CGACGC,CGACGG,CGAGCA,CGAGCG,CGAGGA,CGAGGC,CGAGGG,CGAUGA,CGAUGG,CGCACA,CGCACC,CGCACG,CGCAGC,CGCAGG,CGCCCA,CGCCCC,CGCCCG,CGCCGA,CGCCGC,CGCCGG,CGCCGU,CGCGAG,CGCGCA,CGCGCC,CGCGCG,CGCGCU,CGCGGA,CGCGGC,CGCGGG,CGCGGU,CGCGUC,CGCUAU,CGCUCA,CGCUCC,CGCUCG,CGCUGC,CGCUGG,CGGAAC,CGGAAG,CGGAAU,CGGACA,CGGACC,CGGACG,CGGAGC,CGGAGG,CGGCAC,CGGCAG,CGGCCA,CGGCCC,CGGCCG,CGGCGA,CGGCGC,CGGCGG,CGGGCA,CGGGCC,CGGGCG,CGGGGA,CGGGGC,CGGGGG,CGGUCC,CGGUCG,CGGUGC,CGGUGG,CGUCCA,CGUCCG,CGUCGA,CGUCGG,CGUGCA,CGUGCG,CGUGGA,CGUGGG,CUACGA,CUAGGG,CUCAGG,CUCCGG,CUCGGA,CUCGUG,CUGAAC,CUGACU,CUGAGC,CUGAGU,GAAAGA,GAAAGG,GAACAG,GAACCA,GAACGA,GAAGAA,GAAGAC,GAAGAG,GAAGAU,GAAGCA,GAAGCC,GAAGCU,GAAGGA,GAAGGC,GAAGGU,GAAUGA,GACAGA,GACAGG,GACCCA,GACCCG,GACCGA,GACGAA,GACGAC,GACGAG,GACGCA,GACGCG,GACGGA,GACGGC,GACGGG,GACUGA,GACUGG,GAGAAC,GAGAAG,GAGACA,GAGACG,GAGCAG,GAGCGA,GAGCGG,GAGGAA,GAGGAC,GAGGAG,GAGGAU,GAGGCA,GAGGGA,GAGGGC,GAGGGG,GAGGUA,GAUACG,GAUGAA,GAUGAC,GAUGAU,GAUGCA,GAUGCC,GAUGCU,GAUGGA,GAUGGC,GAUGGU,GCACCA,GCACCG,GCACGA,GCACGG,GCACGU,GCAGCA,GCAGCG,GCAGGA,GCAGGC,GCAGGG,GCAGGU,GCAUAC,GCCCAC,GCCCCA,GCCCCG,GCCCGA,GCCCGG,GCCCGU,GCCGCA,GCCGCG,GCCGGA,GCCGGG,GCCGGU,GCCGUA,GCGAGC,GCGCAA,GCGCCA,GCGCCC,GCGCCG,GCGCGA,GCGCGC,GCGCGG,GCGGAA,GCGGAC,GCGGAG,GCGGCA,GCGGCG,GCGGGA,GCGGGG,GCGGUU,GCGUCA,GCUCCA,GCUCCG,GCUCGA,GCUCGG,GCUGCA,GCUGCG,GCUGGA,GCUGGG,GGAAAG,GGAACA,GGAAGA,GGAAGG,GGAAUG,GGACAA,GGACAG,GGACCA,GGACCG,GGACGA,GGACGG,GGACGU,GGACUG,GGAGAA,GGAGAC,GGAGAU,GGAGCA,GGAGCC,GGAGCG,GGAGCU,GGAGGA,GGAGGC,GGAGGG,GGAGGU,GGAUAC,GGAUAU,GGAUGA,GGAUUC,GGCACA,GGCACG,GGCAGA,GGCCCA,GGCCCG,GGCCGA,GGCCGG,GGCCGU,GGCGCA,GGCGCG,GGCGGA,GGCGGG,GGCGGU,GGGACG,GGGCCA,GGGCCG,GGGCGA,GGGCGG,GGGGAA,GGGGAC,GGGGAG,GGGGCA,GGGGCG,GGGGGA,GGGGGG,GGGUAC,GGUAAC,GGUCCA,GGUCCG,GGUCGA,GGUCGG,GGUGAA,GGUGAC,GGUGAU,GGUGCA,GGUGCC,GGUGCG,GGUGCU,GGUGGA,GGUGGC,GGUGGG,GGUGGU,GUAGGA,GUGACA,GUGACG,UAAUUU,UACGAA,UACGGA,UAGACA,UAGGAC,UAUUAC,UCAAGA,UCAGGU,UCCGGA,UCGCAC,UCGGGC,UCGUGU,UGAACA,UGAACC,UGAAGA,UGAAGC,UGAAGG,UGACAG,UGACGA,UGACUA,UGACUG,UGAGUU,UGAUGA,UGAUGC,UGAUGG,UGGAAA,UGGACA,UGGAGA,UGGAGC,UGGAGG,UGGUGA,UGGUGC,UGGUGG,UGUAGG,UUAAUU,UUACGG,UUCAAG,UUGACA,UUUCAA |
| TRA2B | 2 | 7329 | 0.02127660 | 0.010622665 | 1.002122 | AGGAAA,GAAGGA | AAAGAA,AAGAAC,AAGAAG,AAGAAU,AAGGAA,AAUAAG,AGAAGA,AGAAGG,AGGAAA,AGGAAG,AUAAGA,GAAAGA,GAAGAA,GAAGGA,GAAUUA,GGAAGG,UAAGAA |
RBP on BSJ (Exon Only)
Plot
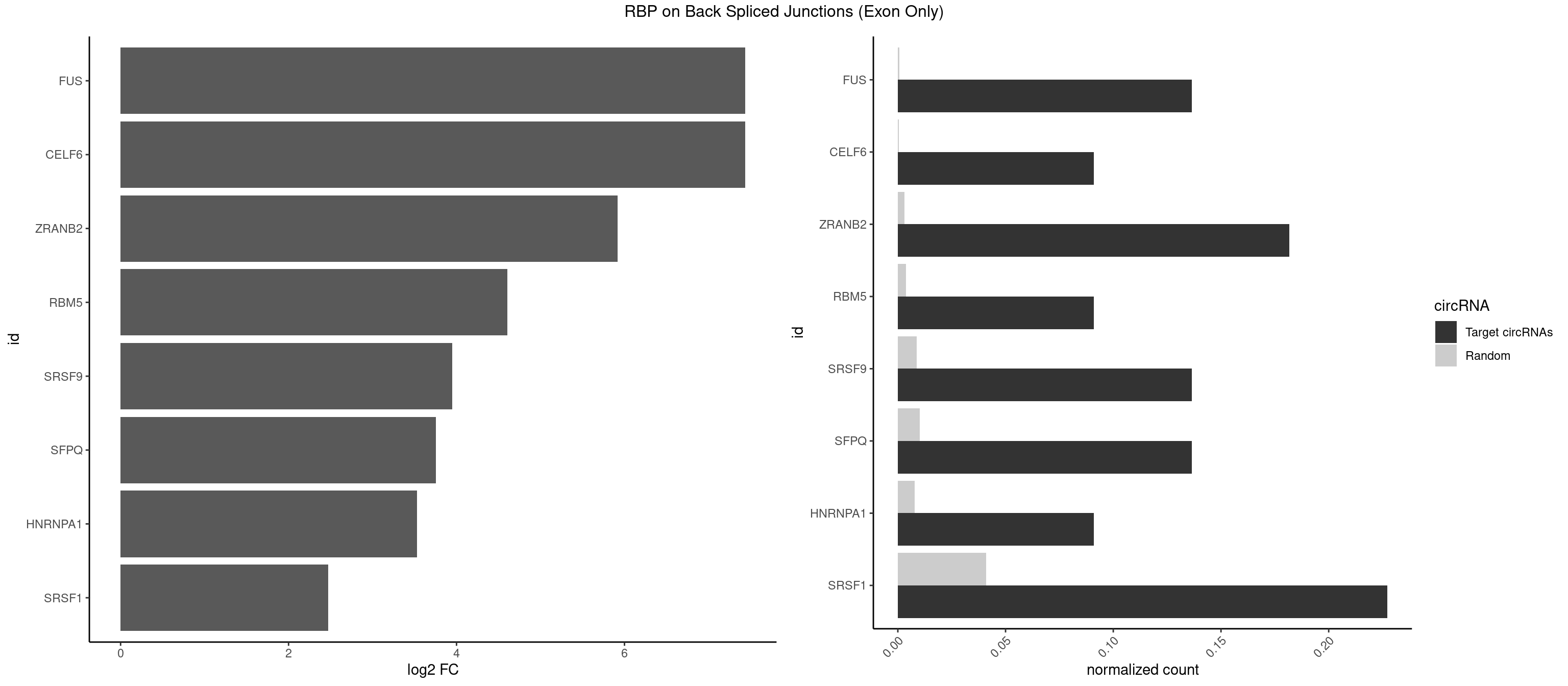
Spreadsheet
| id | foreground | background | foregroundNorm | backgroundNorm | log2FC | motifF | motifB |
|---|---|---|---|---|---|---|---|
| CELF6 | 1 | 7 | 0.09090909 | 0.0005239717 | 7.438792 | GUGGUG | GUGAGG,GUGUUG,UGUGGG,UGUGUG |
| FUS | 2 | 11 | 0.13636364 | 0.0007859576 | 7.438792 | CGGUGG,UGGUGG | AAAAAA,CGGUGG,GGGUGA,GGGUGG,GGGUGU |
| ZRANB2 | 3 | 45 | 0.18181818 | 0.0030128373 | 5.915230 | GGUGGU,GUGGUG,UGGUGG | AAAGGU,AGGGAA,AGGUAA,AGGUAC,AGGUAG,AGGUAU,AGGUCU,AGGUUA,AGGUUU,GAGGUU,GGGUAA,GGGUGA,GGUAAA,GGUGGU,GUAAAG,UAAAGG,UGGUAA |
| RBM5 | 1 | 56 | 0.09090909 | 0.0037332984 | 4.605902 | GGUGGU | AAAAAA,AAGGAA,AAGGGG,AAGGUG,AGGGAA,AGGGAG,AGGGUA,AGGGUG,AGGUAA,CAAGGA,CAAGGG,CUCUUC,GAAGGA,GAAGGG,GAAGGU,GAGGGA,GAGGGU,GGUGGU,UCUUCU |
| SRSF9 | 2 | 134 | 0.13636364 | 0.0088420225 | 3.946939 | GGUGGC,UGGUGG | AGGAAA,AGGAAC,AGGACA,AGGACC,AGGAGA,AGGAGC,AUGAAC,AUGACA,AUGAGA,AUGAGC,CUGGAU,GAAGCA,GAAGCC,GAAGGA,GAAGGC,GAUGCA,GAUGCC,GAUGGA,GAUGGC,GGAAAA,GGAAAG,GGAACA,GGAACG,GGAAGC,GGAAGG,GGACAA,GGACAG,GGACCA,GGACCG,GGAGAA,GGAGAG,GGAGCA,GGAGCG,GGAGGA,GGAGGC,GGAUGG,GGGAGC,GGGUGG,GGUGCA,GGUGCC,GGUGGA,GGUGGC,UGAAAA,UGAACA,UGAACG,UGAAGC,UGAAGG,UGACAA,UGACAG,UGAGAA,UGAGAG,UGAGCA,UGAUGG,UGGAGC,UGGAGG,UGGUGC |
| SFPQ | 2 | 153 | 0.13636364 | 0.0100864553 | 3.756968 | GUGGUG,UGGUGG | AAGAAC,AAGAGC,AAGAGG,AAGCAA,AAGGAA,AAGGAC,AAGGGA,ACUGGG,AGAGAG,AGAGGA,AGAGGU,AGGAAC,AGGACC,AGGGAU,AGGGGG,AUCGGA,CAGGCA,CUGGAG,CUGGGA,GAAGAA,GAAGAG,GAAGCA,GAAGGA,GAGGAA,GAGGAC,GAGGUA,GCAGGC,GGAAGA,GGAGAG,GGAGGA,GGAGGG,GGGGGA,GUAAGA,GUAAUG,GUAGUG,GUAGUU,GUCUGG,GUGAUU,UAAGAG,UAAGGA,UAAGGG,UAAUGG,UAAUUG,UAGAGA,UAGAUC,UAGUGG,UAGUGU,UAGUUG,UCGGAA,UCUAAG,UGAAGC,UGAUGG,UGAUGU,UGAUUG,UGCAGG,UGGAGA,UGGAGC,UGGAGG,UGGUUU,UUAAUG,UUAGUG,UUAGUU,UUGAAG,UUGGUU |
| HNRNPA1 | 1 | 119 | 0.09090909 | 0.0078595756 | 3.531901 | GUGGUG | AAAAAA,AAAGAG,AAGAGG,AAGGUG,AAUUUA,AGAGGA,AGAUAU,AGAUUA,AGGAAG,AGGAGC,AGGGAC,AGGGCA,AGGGUU,AGUAGG,AGUGAA,AUAGGG,AUUAGA,AUUUAA,CAAAGA,CAAGGA,CAGGGA,CCAAGG,GAAGGU,GACUUA,GAGGAA,GAGGAG,GAGUGG,GAUUAG,GCAGGC,GCCAAG,GGAAGG,GGACUU,GGAGGA,GGCAGG,GGGACU,GGGCAG,GGUGCG,GUUAGG,UAGACA,UAGAGA,UAGAGU,UAGAUU,UAGGAA,UAGGCU,UAGGGC,UAGUGA,UAGUUA,UCGGGC,UGGUGC,UUAGAU,UUAGGG,UUUAGA |
| SRSF1 | 4 | 625 | 0.22727273 | 0.0410007860 | 2.470701 | CGGUGG,GGUGGC,GGUGGU,UGGUGG | AAAAGA,AAAGAA,AAAGAC,AAAGAG,AACAGC,AAGAAC,AAGACC,AAGAGA,AAGAGC,AAGAGG,AAGGAC,AAGGCG,AAGGUG,AAUGAC,AAUUUC,ACAAGG,ACAGAG,ACAGCA,ACAGCG,ACAGGA,ACAGGG,ACAGGU,ACAGUG,ACCCGA,ACCGGA,ACGAAU,ACGGAA,ACUGAG,ACUGGA,AGAAGA,AGAAGG,AGACAA,AGACAG,AGACGU,AGAGAA,AGAGAC,AGAGCA,AGAGGA,AGAGGG,AGAGGU,AGAUGG,AGCAGG,AGCCGA,AGCGGA,AGGAAA,AGGAAC,AGGAAG,AGGACA,AGGACC,AGGACG,AGGACU,AGGAGA,AGGAGC,AGGAGG,AGGUAA,AUGAAC,AUGAAG,AUGACA,AUGACU,AUGGAC,AUGGAG,CAAGGA,CAAUGG,CACAGA,CACAGC,CACAGG,CACAGU,CACCCA,CACCCG,CACCGG,CACGCA,CACGGA,CACUGG,CAGAAC,CAGACA,CAGACG,CAGAGA,CAGAGC,CAGAGG,CAGCCG,CAGUCG,CAUGGU,CCAACC,CCAAGG,CCACCA,CCACCC,CCACGG,CCAGCA,CCAGCC,CCAGCG,CCAGGA,CCAGGG,CCCACC,CCCAGC,CCCAGG,CCCCGC,CCCGGG,CCCGUU,CCCUCC,CCCUCG,CCGAGG,CCGCGA,CCGCUA,CCGGAC,CCGGAG,CCGGGA,CCGUCC,CCGUGC,CCGUGG,CCGUUU,CCUAGG,CCUCCG,CCUCGA,CCUGCG,CCUGGA,CCUGGG,CGAACG,CGAAGC,CGAGGA,CGAGGC,CGAGGG,CGAUGG,CGCAGC,CGCCGC,CGCUAU,CGCUGC,CGGAAU,CGGACA,CGGAGC,CGGAGG,CGGCGG,CGGGCA,CGGUGC,CGGUGG,CGUGCG,CGUGGA,CGUGGG,CUCAGG,CUCGUG,CUGAAC,CUGAGU,GAAAGA,GAAAGG,GAACAG,GAAGAA,GAAGAG,GAAGAU,GAAGCA,GAAGCC,GAAGCU,GAAGGA,GAAGGC,GAAGGU,GAAUGA,GACAGA,GACAGG,GACCCA,GACGAA,GACGAC,GACGGA,GACUGA,GAGAAC,GAGAAG,GAGACA,GAGACG,GAGCAG,GAGGAA,GAGGAC,GAGGAG,GAGGAU,GAGGCA,GAGGGA,GAGGGC,GAGGGG,GAGGUA,GAUGAA,GAUGAC,GAUGAU,GAUGCA,GAUGCC,GAUGCU,GAUGGA,GAUGGC,GCACGG,GCAGCA,GCAGCG,GCAGGA,GCAGGC,GCAGGG,GCAGGU,GCCCAC,GCCCGG,GCCCGU,GCGCAA,GCGCCA,GCGCCC,GCGCGG,GCGGAC,GCGGCG,GCGGUU,GCUGGG,GGAAAG,GGAACA,GGAAGA,GGAAGG,GGAAUG,GGACAA,GGACAG,GGACCA,GGACCG,GGACGA,GGAGAA,GGAGAC,GGAGAU,GGAGCA,GGAGCG,GGAGCU,GGAGGA,GGAGGC,GGAGGG,GGAGGU,GGAUAU,GGAUGA,GGAUUC,GGCACA,GGCAGA,GGCCGA,GGCGCA,GGGACG,GGGCCG,GGGGAA,GGGGAG,GGGGCA,GGGGCG,GGGGGA,GGGUAC,GGUCCA,GGUCCG,GGUGAA,GGUGCA,GGUGCC,GGUGCG,GGUGCU,GGUGGA,GGUGGC,GGUGGG,GGUGGU,GUAGGA,GUGACA,UAGACA,UAGGAC,UCAAGA,UCAGGU,UCGGGC,UGAACA,UGAAGA,UGAAGC,UGAAGG,UGACAG,UGACGA,UGACUG,UGAGUU,UGAUGA,UGAUGG,UGGACA,UGGAGA,UGGAGC,UGGAGG,UGGUGC,UGUAGG,UUCAAG |
RBP on BSJ (Exon and Intron)
Plot
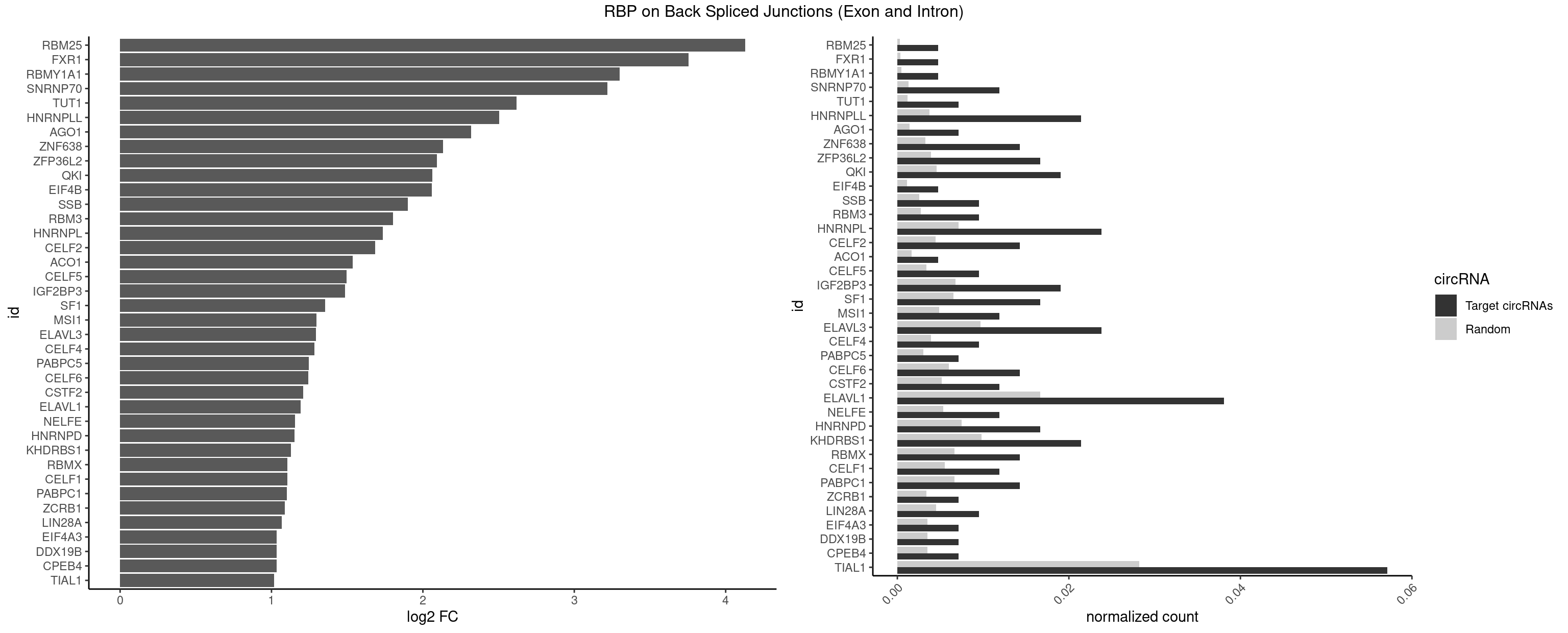
Spreadsheet
| id | foreground | background | foregroundNorm | backgroundNorm | log2FC | motifF | motifB |
|---|---|---|---|---|---|---|---|
| RBM25 | 1 | 53 | 0.004761905 | 0.0002720677 | 4.129501 | CGGGCA | AUCGGG,CGGGCA,UCGGGC |
| FXR1 | 1 | 69 | 0.004761905 | 0.0003526804 | 3.755106 | AUGACA | ACGACA,ACGACG,AUGACA,AUGACG |
| RBMY1A1 | 1 | 95 | 0.004761905 | 0.0004836759 | 3.299426 | CAAGAC | ACAAGA,CAAGAC |
| SNRNP70 | 4 | 253 | 0.011904762 | 0.0012797259 | 3.217632 | GAUCAA,GUUCAA,UUCAAG | AAUCAA,AUCAAG,AUUCAA,GAUCAA,GUUCAA,UUCAAG |
| TUT1 | 2 | 230 | 0.007142857 | 0.0011638452 | 2.617602 | AAAUAC,AAUACU | AAAUAC,AAUACU,AGAUAC,CAAUAC,CGAUAC,GAUACU |
| HNRNPLL | 8 | 748 | 0.021428571 | 0.0037736800 | 2.505492 | ACACAC,ACACCA,CACACA,CACACC,CACCAC | ACAAAC,ACACAC,ACACCA,ACAUAC,ACCACA,ACCGCA,ACGACA,ACUGCA,AGACGA,CAAACA,CACACA,CACACC,CACCAC,CACCGC,CACUGC,CAGACG,CAUACA,GACGAC,GCAAAC,GCACAC,GCAUAC |
| AGO1 | 2 | 283 | 0.007142857 | 0.0014308746 | 2.319604 | GGUAGU,UGAGGU | AGGUAG,GAGGUA,GGUAGU,GUAGUA,UGAGGU |
| ZNF638 | 5 | 646 | 0.014285714 | 0.0032597743 | 2.131729 | GGUUGG,GUUGGU,GUUGUU,UGUUCU,UGUUGU | CGUUCG,CGUUCU,CGUUGG,CGUUGU,GGUUCG,GGUUCU,GGUUGG,GGUUGU,GUUCGU,GUUCUU,GUUGGU,GUUGUU,UGUUCG,UGUUCU,UGUUGG,UGUUGU |
| ZFP36L2 | 6 | 774 | 0.016666667 | 0.0039046755 | 2.093691 | UAUUUA,UUAUUU,UUUAUU | AUUUAU,UAUUUA,UUAUUU,UUUAUU |
| QKI | 7 | 904 | 0.019047619 | 0.0045596534 | 2.062615 | ACACAC,CUAACA,CUACUC,UACUCA | AAUCAU,ACACAC,ACACUA,ACUAAC,ACUAAU,ACUCAU,ACUUAU,AUCAUA,AUCUAA,AUCUAC,AUUAAC,CACACU,CACUAA,CUAACA,CUAACC,CUAACG,CUAAUC,CUACUC,CUCAUA,UAACCU,UAAUCA,UACUCA,UCAUAU,UCUAAU,UCUACU |
| EIF4B | 1 | 226 | 0.004761905 | 0.0011436921 | 2.057840 | UUGGAA | CUCGGA,CUUGGA,GUCGGA,GUUGGA,UCGGAA,UCGGAC,UUGGAA,UUGGAC |
| SSB | 3 | 505 | 0.009523810 | 0.0025493753 | 1.901395 | UGCUGU,UGUUUU | CUGUUU,GCUGUU,UGCUGU,UGUUUU |
| RBM3 | 3 | 541 | 0.009523810 | 0.0027307537 | 1.802240 | AAUACU,AUACUA,GAGACU | AAAACG,AAAACU,AAACGA,AAACUA,AAGACG,AAGACU,AAUACG,AAUACU,AGACGA,AGACUA,AUACGA,AUACUA,GAAACG,GAAACU,GAGACG,GAGACU,GAUACG,GAUACU |
| HNRNPL | 9 | 1419 | 0.023809524 | 0.0071543732 | 1.734641 | AAAUAC,ACACAC,ACACCA,CACACA,CACACC,CACCAC | AAACAA,AAACAC,AAAUAA,AAAUAC,AACAAA,AACACA,AAUAAA,AAUACA,ACACAA,ACACAC,ACACCA,ACACGA,ACAUAA,ACAUAC,ACCACA,CACAAA,CACAAC,CACAAG,CACACA,CACACC,CACCAC,CACGAA,CACGAC,CACGAG,CAUAAA,CAUACA |
| CELF2 | 5 | 881 | 0.014285714 | 0.0044437727 | 1.684716 | GUUGUU,UAUGUU,UGUGUG,UGUUGU,UUGUGU | AUGUGU,GUAUGU,GUCUGU,GUGUGU,GUUGUU,UAUGUG,UAUGUU,UGUGUG,UGUUGU,UUGUGU |
| ACO1 | 1 | 325 | 0.004761905 | 0.0016424829 | 1.535660 | CAGUGU | CAGUGA,CAGUGC,CAGUGG,CAGUGU |
| CELF5 | 3 | 669 | 0.009523810 | 0.0033756550 | 1.496371 | GUGUGG,GUGUUG,UGUGUG | GUGUGG,GUGUGU,GUGUUG,GUGUUU,UGUGUG,UGUGUU |
| IGF2BP3 | 7 | 1348 | 0.019047619 | 0.0067966546 | 1.486714 | AAAUAC,ACACAC,CACACA,CAUUCA | AAAAAC,AAAACA,AAAAUC,AAACAC,AAACUC,AAAUAC,AAAUCA,AAAUUC,AACACA,AACUCA,AAUACA,AAUUCA,ACAAAC,ACAAUC,ACACAC,ACACUC,ACAUAC,ACAUUC,CAAACA,CAAUCA,CACACA,CACUCA,CAUACA,CAUUCA |
| SF1 | 6 | 1294 | 0.016666667 | 0.0065245869 | 1.353007 | AUACUA,CUAACA,GCUAAC,UACUAA,UAGUAA,UGCUAA | ACAGAC,ACAGUC,ACCGAC,ACGAAC,ACUAAC,ACUAAG,ACUAAU,ACUAGC,ACUGAC,ACUUAU,AGUAAC,AGUAAG,AUACUA,AUUAAC,CACAGA,CACCGA,CACUGA,CAGUCA,CGCUGA,CUAACA,GACUAA,GCUAAC,GCUGAC,GCUGCC,UAACAA,UACGAA,UACUAA,UACUAG,UACUGA,UAGUAA,UAUACU,UGCUAA,UGCUGA,UGCUGC |
| MSI1 | 4 | 961 | 0.011904762 | 0.0048468360 | 1.296424 | AGGUGG,UAGGUG,UAGUAA | AGGAAG,AGGAGG,AGGUAG,AGGUGG,AGUAAG,AGUAGG,AGUUAG,AGUUGG,UAGGAA,UAGGAG,UAGGUA,UAGGUG,UAGUAA,UAGUAG,UAGUUA,UAGUUG |
| ELAVL3 | 9 | 1928 | 0.023809524 | 0.0097188634 | 1.292679 | AUUUUA,UAUUUA,UUAUUU,UUUAUU,UUUUUU | AUUUAU,AUUUUA,UAUUUA,UAUUUU,UUAUUU,UUUAUU,UUUUUU |
| CELF4 | 3 | 776 | 0.009523810 | 0.0039147521 | 1.282618 | GUGUGG,GUGUUG,UGUGUG | GGUGUG,GGUGUU,GUGUGG,GUGUGU,GUGUUG,GUGUUU,UGUGUG,UGUGUU |
| PABPC5 | 2 | 596 | 0.007142857 | 0.0030078597 | 1.247764 | AGAAAA,GAAAGU | AGAAAA,AGAAAG,AGAAAU,GAAAAU,GAAAGU,GAAAUU |
| CELF6 | 5 | 1196 | 0.014285714 | 0.0060308343 | 1.244144 | GUGUGG,GUGUUG,UGUGGU,UGUGUG | GUGAGG,GUGAUG,GUGGGG,GUGGUG,GUGUGG,GUGUUG,UGUGAG,UGUGAU,UGUGGG,UGUGGU,UGUGUG,UGUGUU |
| CSTF2 | 4 | 1022 | 0.011904762 | 0.0051541717 | 1.207726 | GUGUUG,UGUGUG,UGUUUU | GUGUGU,GUGUUG,GUGUUU,GUUUUG,UGUGUG,UGUGUU,UGUUUG,UGUUUU |
| ELAVL1 | 15 | 3309 | 0.038095238 | 0.0166767432 | 1.191773 | UAUUUA,UGUUUU,UUAUUU,UUGUUU,UUUAGU,UUUAUU,UUUGUU,UUUUUG,UUUUUU | AUUUAU,CCCCCC,UAGUUU,UAUUUA,UAUUUU,UGAUUU,UGGUUU,UGUUUU,UUAGUU,UUAUUU,UUGAUU,UUGGUU,UUGUUU,UUUAGU,UUUAUU,UUUGGU,UUUGUU,UUUUUG,UUUUUU |
| NELFE | 4 | 1059 | 0.011904762 | 0.0053405885 | 1.156468 | CUGGUU,GCUAAC,GUCUCU,UGGUUA | CUCUCU,CUCUGG,CUGGCU,CUGGUU,GCUAAC,GGCUAA,GGUCUC,GGUUAG,GUCUCU,UCUCUC,UCUCUG,UCUGGC,UCUGGU,UGGCUA,UGGUUA |
| HNRNPD | 6 | 1488 | 0.016666667 | 0.0075020153 | 1.151615 | UAUUUA,UUAUUU,UUUAUU | AAAAAA,AAUUUA,AGAUAU,AGUAGG,AUUUAU,UAUUUA,UUAGAG,UUAGGA,UUAGGG,UUAUUU,UUUAUU |
| KHDRBS1 | 8 | 1945 | 0.021428571 | 0.0098045143 | 1.128018 | AUUUAA,CUAAAA,UAAAAG,UUAAAA,UUAAAU,UUUUUU | AUAAAA,AUUUAA,AUUUAC,CAAAAU,CUAAAA,CUAAAU,GAAAAC,UAAAAA,UAAAAC,UAAAAG,UUAAAA,UUAAAU,UUUUAC,UUUUUU |
| RBMX | 5 | 1316 | 0.014285714 | 0.0066354293 | 1.106311 | AAGUGU,ACCAAA,AGUGUU,AUCAAA | AAGAAG,AAGGAA,AAGUAA,AAGUGU,ACCAAA,AGAAGG,AGGAAG,AGUAAC,AGUGUU,AUCAAA,AUCCCA,AUCCCC,GAAGGA,GGAAGG,GUAACA,UAACAA,UAAGAC,UCAAAA |
| CELF1 | 4 | 1097 | 0.011904762 | 0.0055320435 | 1.105654 | GUGUUG,UGUGUG,UGUUGU,UUGUGU | CUGUCU,GUGUGU,GUGUUG,GUUGUG,GUUUGU,UGUCUG,UGUGUG,UGUGUU,UGUUGU,UGUUUG,UUGUGU |
| PABPC1 | 5 | 1321 | 0.014285714 | 0.0066606207 | 1.100845 | ACAAAU,AGAAAA,CAAACC,CUAACA,GAAAAA | AAAAAA,AAAAAC,ACAAAC,ACAAAU,ACGAAC,ACGAAU,ACUAAC,ACUAAU,AGAAAA,CAAACA,CAAACC,CAAAUA,CAAAUC,CGAACA,CGAACC,CGAAUA,CGAAUC,CUAACA,CUAACC,CUAAUA,CUAAUC,GAAAAA,GAAAAC |
| ZCRB1 | 2 | 666 | 0.007142857 | 0.0033605401 | 1.087808 | ACUUAA,AUUUAA | AAUUAA,ACUUAA,AUUUAA,GAAUUA,GACUUA,GAUUAA,GAUUUA,GCUUAA,GGAUUA,GGCUUA,GGUUUA,GUUUAA |
| LIN28A | 3 | 900 | 0.009523810 | 0.0045395002 | 1.069005 | AGGAGA,AGGAGU,GGAGAA | AGGAGA,AGGAGU,CGGAGA,CGGAGG,CGGAGU,GGAGAA,GGAGAU,GGAGGA,GGAGGG,GGAGUA,UGGAGA,UGGAGG,UGGAGU |
| CPEB4 | 2 | 691 | 0.007142857 | 0.0034864974 | 1.034723 | UUUUUU | UUUUUU |
| DDX19B | 2 | 691 | 0.007142857 | 0.0034864974 | 1.034723 | UUUUUU | UUUUUU |
| EIF4A3 | 2 | 691 | 0.007142857 | 0.0034864974 | 1.034723 | UUUUUU | UUUUUU |
| TIAL1 | 23 | 5597 | 0.057142857 | 0.0282043531 | 1.018655 | AAAUUU,AAUUUU,AGUUUU,AUUUUA,AUUUUC,GUUUUU,UUAAAU,UUAUUU,UUCAGU,UUUAAA,UUUAUU,UUUUAU,UUUUUA,UUUUUG,UUUUUU | AAAUUU,AAUUUU,AGUUUU,AUUUUA,AUUUUC,AUUUUG,AUUUUU,CAGUUU,CUUUUA,CUUUUC,CUUUUG,CUUUUU,GUUUUC,GUUUUU,UAAAUU,UAUUUU,UCAGUU,UUAAAU,UUAUUU,UUCAGU,UUUAAA,UUUAUU,UUUCAG,UUUUAA,UUUUAU,UUUUCA,UUUUUA,UUUUUC,UUUUUG,UUUUUU |
Help
- Note:
- foreground: number of motifs found in the foreground target sequences (e.g. predicted circRNAs).
- background: number of motifs found in the background target sequences (e.g. randomly generated circRNAs).
- foregroundNorm: number of motifs found in the foreground target sequences (+1) divided by the number (or length) of sequences analyzed.
- backgroundNorm: number of motifs found in background target sequences (+1) divided by the number (or length) of sequences analyzed.
- log2FC: log2 fold change calculated in this way (foregroundNorm)/(backgroundNorm).
- motifF: motifs of the corresponding RBP found in the foreground sequences.
- motifB: motifs of the corresponding RBP found in the background sequences.