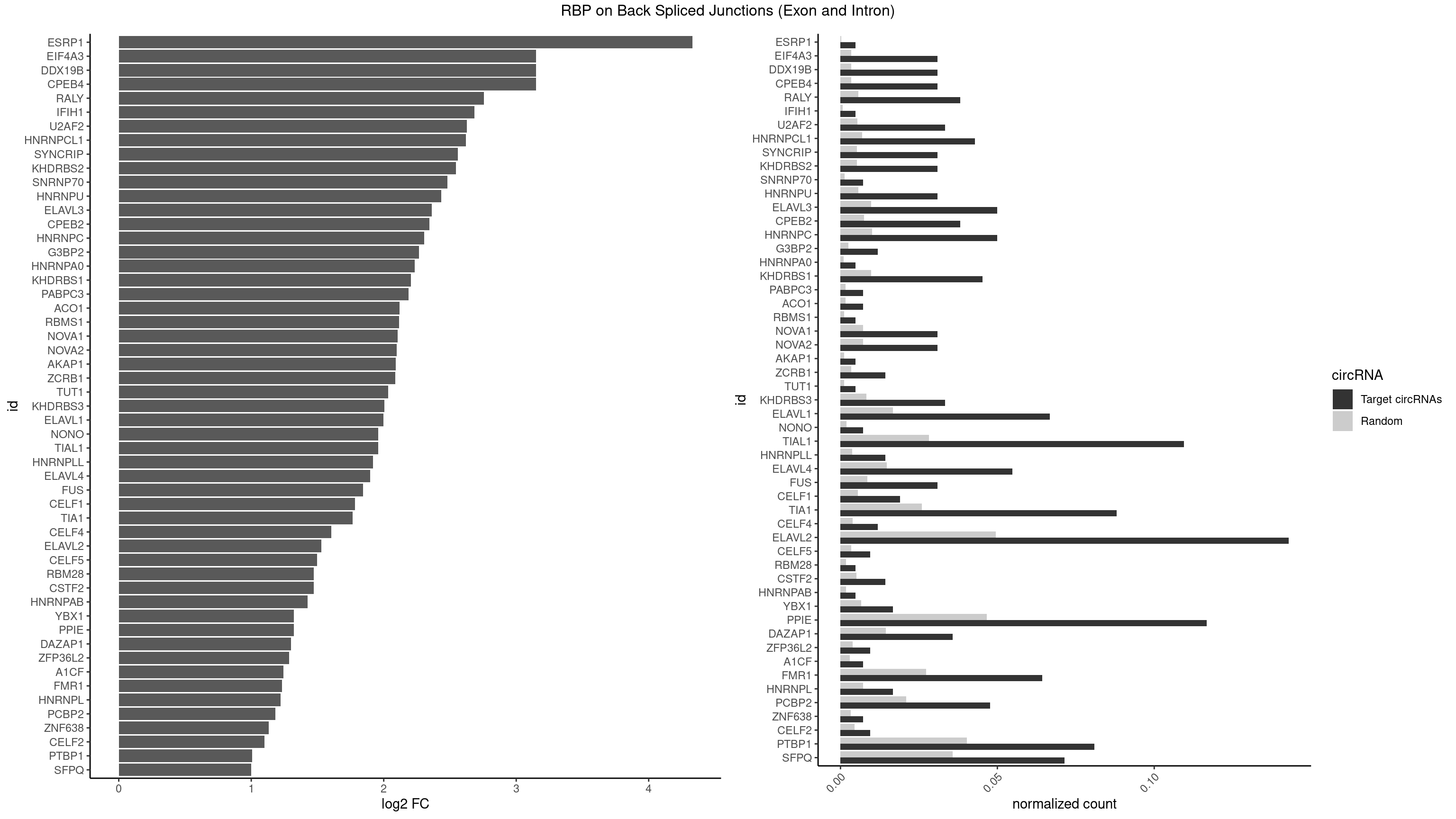
| ESRP1 |
1 |
46 |
0.004761905 |
0.0002367997 |
4.329800 |
AGGGAU |
AGGGAU |
| CPEB4 |
12 |
691 |
0.030952381 |
0.0034864974 |
3.150200 |
UUUUUU |
UUUUUU |
| DDX19B |
12 |
691 |
0.030952381 |
0.0034864974 |
3.150200 |
UUUUUU |
UUUUUU |
| EIF4A3 |
12 |
691 |
0.030952381 |
0.0034864974 |
3.150200 |
UUUUUU |
UUUUUU |
| RALY |
15 |
1117 |
0.038095238 |
0.0056328094 |
2.757684 |
UUUUUC,UUUUUG,UUUUUU |
UUUUUC,UUUUUG,UUUUUU |
| IFIH1 |
1 |
146 |
0.004761905 |
0.0007406288 |
2.684716 |
GGCCCU |
CCGCGG,CGCGGA,GCCGCG,GCGGAU,GGCCCU,GGCCGC,GGGCCG |
| U2AF2 |
13 |
1071 |
0.033333333 |
0.0054010480 |
2.625654 |
UUUUUC,UUUUUU |
UUUUCC,UUUUUC,UUUUUU |
| HNRNPCL1 |
17 |
1381 |
0.042857143 |
0.0069629182 |
2.621772 |
AUUUUU,CUUUUU,UUUUUG,UUUUUU |
AUUUUU,CUUUUU,UUUUUG,UUUUUU |
| SYNCRIP |
12 |
1041 |
0.030952381 |
0.0052498992 |
2.559689 |
UUUUUU |
AAAAAA,UUUUUU |
| KHDRBS2 |
12 |
1051 |
0.030952381 |
0.0053002821 |
2.545909 |
UUUUUU |
AAUAAA,AUAAAA,AUAAAC,GAUAAA,UUUUUU |
| SNRNP70 |
2 |
253 |
0.007142857 |
0.0012797259 |
2.480666 |
GUUCAA,UUCAAG |
AAUCAA,AUCAAG,AUUCAA,GAUCAA,GUUCAA,UUCAAG |
| HNRNPU |
12 |
1136 |
0.030952381 |
0.0057285369 |
2.433812 |
UUUUUU |
AAAAAA,CCCCCC,GGGGGG,UUUUUU |
| ELAVL3 |
20 |
1928 |
0.050000000 |
0.0097188634 |
2.363069 |
AUUUUA,UAUUUA,UAUUUU,UUAUUU,UUUUUU |
AUUUAU,AUUUUA,UAUUUA,UAUUUU,UUAUUU,UUUAUU,UUUUUU |
| CPEB2 |
15 |
1487 |
0.038095238 |
0.0074969770 |
2.345230 |
AUUUUU,CUUUUU,UUUUUU |
AUUUUU,CAUUUU,CCUUUU,CUUUUU,UUUUUU |
| HNRNPC |
20 |
2006 |
0.050000000 |
0.0101118501 |
2.305881 |
AUUUUU,CUUUUU,UUUUUA,UUUUUC,UUUUUG,UUUUUU |
AUUUUU,CUUUUU,GCAUAC,GGAUAC,GGAUAU,GGAUUC,GGGGGG,GGGUAC,UUUUUA,UUUUUC,UUUUUG,UUUUUU |
| G3BP2 |
4 |
490 |
0.011904762 |
0.0024738009 |
2.266737 |
AGGAUU,GGAUGA,GGAUUA,GGAUUG |
AGGAUA,AGGAUG,AGGAUU,GGAUAA,GGAUAG,GGAUGA,GGAUGG,GGAUUA,GGAUUG |
| HNRNPA0 |
1 |
200 |
0.004761905 |
0.0010126965 |
2.233337 |
AGAUAU |
AAUUUA,AGAUAU,AGUAGG |
| KHDRBS1 |
18 |
1945 |
0.045238095 |
0.0098045143 |
2.206020 |
AUUUAA,AUUUAC,UAAAAA,UUAAAA,UUAAAU,UUUUAC,UUUUUU |
AUAAAA,AUUUAA,AUUUAC,CAAAAU,CUAAAA,CUAAAU,GAAAAC,UAAAAA,UAAAAC,UAAAAG,UUAAAA,UUAAAU,UUUUAC,UUUUUU |
| PABPC3 |
2 |
310 |
0.007142857 |
0.0015669085 |
2.188580 |
AAAAAC,AAAACA |
AAAAAC,AAAACA,AAAACC,GAAAAC |
| ACO1 |
2 |
325 |
0.007142857 |
0.0016424829 |
2.120623 |
CAGUGG,CAGUGU |
CAGUGA,CAGUGC,CAGUGG,CAGUGU |
| RBMS1 |
1 |
217 |
0.004761905 |
0.0010983474 |
2.116204 |
GAUAUA |
AUAUAC,AUAUAG,GAUAUA,UAUAUA |
| NOVA1 |
12 |
1428 |
0.030952381 |
0.0071997179 |
2.104038 |
UUUUUU |
AACACC,ACCACC,AGCACC,AGUCAC,AUCAAC,AUCACC,AUCAUC,AUUCAU,CAGUCA,CAUUCA,CCCCCC,GGGGGG,UCAGUC,UCAUUC,UUCAUU,UUUUUU |
| NOVA2 |
12 |
1434 |
0.030952381 |
0.0072299476 |
2.097993 |
UUUUUU |
AACACC,ACCACC,AGACAU,AGAUCA,AGCACC,AGGCAU,AGUCAU,AUCAAC,AUCACC,AUCAUC,CCUAGA,CUAGAU,GAGACA,GAGGCA,GAGUCA,GAUCAC,GGGGGG,UAGAUC,UUUUUU |
| AKAP1 |
1 |
221 |
0.004761905 |
0.0011185006 |
2.089973 |
AUAUAU |
AUAUAU,UAUAUA |
| ZCRB1 |
5 |
666 |
0.014285714 |
0.0033605401 |
2.087808 |
AUUUAA,GAAUUA,GAUUAA,GAUUUA,GGAUUA |
AAUUAA,ACUUAA,AUUUAA,GAAUUA,GACUUA,GAUUAA,GAUUUA,GCUUAA,GGAUUA,GGCUUA,GGUUUA,GUUUAA |
| TUT1 |
1 |
230 |
0.004761905 |
0.0011638452 |
2.032640 |
AAUACU |
AAAUAC,AAUACU,AGAUAC,CAAUAC,CGAUAC,GAUACU |
| KHDRBS3 |
13 |
1646 |
0.033333333 |
0.0082980653 |
2.006119 |
AUUAAA,UUUUUU |
AAAUAA,AAUAAA,AGAUAA,AUAAAA,AUAAAC,AUAAAG,AUUAAA,GAUAAA,UAAAAC,UAAAUA,UUAAAC,UUUUUU |
| ELAVL1 |
27 |
3309 |
0.066666667 |
0.0166767432 |
1.999128 |
UAUUUA,UAUUUU,UGAUUU,UGGUUU,UGUUUU,UUAUUU,UUGUUU,UUUGUU,UUUUUG,UUUUUU |
AUUUAU,CCCCCC,UAGUUU,UAUUUA,UAUUUU,UGAUUU,UGGUUU,UGUUUU,UUAGUU,UUAUUU,UUGAUU,UUGGUU,UUGUUU,UUUAGU,UUUAUU,UUUGGU,UUUGUU,UUUUUG,UUUUUU |
| NONO |
2 |
364 |
0.007142857 |
0.0018389762 |
1.957598 |
AGAGGA,GAGGAA |
AGAGGA,AGGAAC,GAGAGG,GAGGAA |
| TIAL1 |
45 |
5597 |
0.109523810 |
0.0282043531 |
1.957255 |
AAAUUU,AAUUUU,AUUUUA,AUUUUC,AUUUUG,AUUUUU,CAGUUU,CUUUUG,CUUUUU,GUUUUU,UAAAUU,UAUUUU,UCAGUU,UUAAAU,UUAUUU,UUCAGU,UUUAAA,UUUUAA,UUUUUA,UUUUUC,UUUUUG,UUUUUU |
AAAUUU,AAUUUU,AGUUUU,AUUUUA,AUUUUC,AUUUUG,AUUUUU,CAGUUU,CUUUUA,CUUUUC,CUUUUG,CUUUUU,GUUUUC,GUUUUU,UAAAUU,UAUUUU,UCAGUU,UUAAAU,UUAUUU,UUCAGU,UUUAAA,UUUAUU,UUUCAG,UUUUAA,UUUUAU,UUUUCA,UUUUUA,UUUUUC,UUUUUG,UUUUUU |
| HNRNPLL |
5 |
748 |
0.014285714 |
0.0037736800 |
1.920529 |
ACCACA,ACUGCA,CACACC,CACCAC,CACUGC |
ACAAAC,ACACAC,ACACCA,ACAUAC,ACCACA,ACCGCA,ACGACA,ACUGCA,AGACGA,CAAACA,CACACA,CACACC,CACCAC,CACCGC,CACUGC,CAGACG,CAUACA,GACGAC,GCAAAC,GCACAC,GCAUAC |
| ELAVL4 |
22 |
2916 |
0.054761905 |
0.0146966949 |
1.897681 |
UAUUUA,UAUUUU,UUAUUU,UUGUAU,UUUGUA,UUUUUA,UUUUUU |
AAAAAA,AUCUAA,AUUUAU,GUAUCU,UAUCUA,UAUUUA,UAUUUU,UCUAAU,UGUAUC,UUAUUU,UUGUAU,UUUAUU,UUUGUA,UUUUAU,UUUUUA,UUUUUU |
| FUS |
12 |
1711 |
0.030952381 |
0.0086255542 |
1.843361 |
UUUUUU |
AAAAAA,CGGUGA,CGGUGG,GGGGGG,GGGUGA,GGGUGC,GGGUGG,GGGUGU,UGGUGA,UGGUGG,UGGUGU,UUUUUU |
| CELF1 |
7 |
1097 |
0.019047619 |
0.0055320435 |
1.783726 |
CUGUCU,GUGUGU,GUGUUG,GUUUGU,UGUGUG,UGUUGU,UGUUUG |
CUGUCU,GUGUGU,GUGUUG,GUUGUG,GUUUGU,UGUCUG,UGUGUG,UGUGUU,UGUUGU,UGUUUG,UUGUGU |
| TIA1 |
36 |
5140 |
0.088095238 |
0.0259018541 |
1.766009 |
AUUUUC,AUUUUG,AUUUUU,CUUUUG,CUUUUU,GUUUUU,UAUUUU,UCUCCU,UUAUUU,UUCUCC,UUUUCU,UUUUGU,UUUUUA,UUUUUC,UUUUUG,UUUUUU |
AUUUUC,AUUUUG,AUUUUU,CCUUUC,CUCCUU,CUUUUA,CUUUUC,CUUUUG,CUUUUU,GUUUUA,GUUUUC,GUUUUG,GUUUUU,UAUUUU,UCCUUU,UCUCCU,UUAUUU,UUCUCC,UUUAUU,UUUUAU,UUUUCG,UUUUCU,UUUUGG,UUUUGU,UUUUUA,UUUUUC,UUUUUG,UUUUUU |
| CELF4 |
4 |
776 |
0.011904762 |
0.0039147521 |
1.604546 |
GGUGUG,GUGUGU,GUGUUG,UGUGUG |
GGUGUG,GGUGUU,GUGUGG,GUGUGU,GUGUUG,GUGUUU,UGUGUG,UGUGUU |
| ELAVL2 |
59 |
9826 |
0.142857143 |
0.0495112858 |
1.528744 |
AAUUUU,AUUAUU,AUUUAA,AUUUAC,AUUUUA,AUUUUC,AUUUUG,AUUUUU,CUUUUU,GAUUUA,GUUUUU,UAUACU,UAUAUU,UAUUAU,UAUUGA,UAUUUA,UAUUUU,UCUUUU,UGAUUU,UGUAUU,UUAAUU,UUAUAU,UUAUUG,UUAUUU,UUCUUU,UUGUAU,UUUAAG,UUUAAU,UUUACU,UUUAUA,UUUCUU,UUUUAA,UUUUAG,UUUUUA,UUUUUC,UUUUUG,UUUUUU |
AAUUUA,AAUUUG,AAUUUU,AUACUU,AUAUUU,AUUAUU,AUUUAA,AUUUAC,AUUUAG,AUUUAU,AUUUUA,AUUUUC,AUUUUG,AUUUUU,CAUUUU,CUUAUA,CUUUAA,CUUUCU,CUUUUA,CUUUUU,GAUUUA,GAUUUU,GUAUUG,GUUUUA,GUUUUC,GUUUUU,UAAGUU,UAAUUU,UACUUU,UAGUUA,UAUACU,UAUAUA,UAUAUU,UAUGUU,UAUUAU,UAUUGA,UAUUGU,UAUUUA,UAUUUG,UAUUUU,UCAUUU,UCUUAU,UCUUUU,UGAUUU,UGUAUU,UUAAGU,UUAAUU,UUACUU,UUAGUU,UUAUAC,UUAUAU,UUAUGU,UUAUUG,UUAUUU,UUCAUU,UUCUUU,UUGAUU,UUGUAU,UUUAAG,UUUAAU,UUUACU,UUUAGU,UUUAUA,UUUAUG,UUUAUU,UUUCAU,UUUCUU,UUUGAU,UUUUAA,UUUUAG,UUUUAU,UUUUUA,UUUUUC,UUUUUG,UUUUUU |
| CELF5 |
3 |
669 |
0.009523810 |
0.0033756550 |
1.496371 |
GUGUGU,GUGUUG,UGUGUG |
GUGUGG,GUGUGU,GUGUUG,GUGUUU,UGUGUG,UGUGUU |
| RBM28 |
1 |
340 |
0.004761905 |
0.0017180572 |
1.470761 |
GAGUAG |
AGUAGA,AGUAGG,AGUAGU,GAGUAG,GUGUAG,UGUAGA,UGUAGG,UGUAGU |
| CSTF2 |
5 |
1022 |
0.014285714 |
0.0051541717 |
1.470761 |
GUGUGU,GUGUUG,UGUGUG,UGUUUG,UGUUUU |
GUGUGU,GUGUUG,GUGUUU,GUUUUG,UGUGUG,UGUGUU,UGUUUG,UGUUUU |
| HNRNPAB |
1 |
351 |
0.004761905 |
0.0017734784 |
1.424957 |
ACAAAG |
AAAGAC,AAGACA,ACAAAG,AGACAA,AUAGCA,CAAAGA,GACAAA |
| YBX1 |
6 |
1321 |
0.016666667 |
0.0066606207 |
1.323237 |
ACCACA,CACACC,CACCAC,CCACAA,CCACAC,CCACCA |
AACAUC,AACCAC,ACACCA,ACAUCA,ACAUCG,ACAUCU,ACCACA,ACCACC,AUCAUC,CAACCA,CACACC,CACCAC,CAGCAA,CAUCAU,CAUCGC,CAUCUG,CCACAA,CCACAC,CCACCA,CCAGCA,CCCUGC,CCUGCG,CUGCGG,GAUCUG,GCCUGC,GGUCUG,GUCUGC,UCCAGC,UGCGGU |
| PPIE |
48 |
9262 |
0.116666667 |
0.0466696896 |
1.321835 |
AAAUUU,AAUUAU,AAUUUU,AUAUAU,AUAUUA,AUUAAA,AUUAUU,AUUUAA,AUUUUA,AUUUUU,UAAAAA,UAAAUU,UAAUUA,UAUAUU,UAUUAU,UAUUUA,UAUUUU,UUAAAA,UUAAAU,UUAAUU,UUAUAU,UUAUUU,UUUAAA,UUUAAU,UUUAUA,UUUUAA,UUUUUA,UUUUUU |
AAAAAA,AAAAAU,AAAAUA,AAAAUU,AAAUAA,AAAUAU,AAAUUA,AAAUUU,AAUAAA,AAUAAU,AAUAUA,AAUAUU,AAUUAA,AAUUAU,AAUUUA,AAUUUU,AUAAAA,AUAAAU,AUAAUA,AUAAUU,AUAUAA,AUAUAU,AUAUUA,AUAUUU,AUUAAA,AUUAAU,AUUAUA,AUUAUU,AUUUAA,AUUUAU,AUUUUA,AUUUUU,UAAAAA,UAAAAU,UAAAUA,UAAAUU,UAAUAA,UAAUAU,UAAUUA,UAAUUU,UAUAAA,UAUAAU,UAUAUA,UAUAUU,UAUUAA,UAUUAU,UAUUUA,UAUUUU,UUAAAA,UUAAAU,UUAAUA,UUAAUU,UUAUAA,UUAUAU,UUAUUA,UUAUUU,UUUAAA,UUUAAU,UUUAUA,UUUAUU,UUUUAA,UUUUAU,UUUUUA,UUUUUU |
| DAZAP1 |
14 |
2878 |
0.035714286 |
0.0145052398 |
1.299927 |
AGAUAU,AGUUAA,UUUUUU |
AAAAAA,AAUUUA,AGAUAU,AGGAAA,AGGAAG,AGGAUA,AGGAUG,AGGUAA,AGGUAG,AGGUUA,AGGUUG,AGUAAA,AGUAAG,AGUAGG,AGUAUA,AGUAUG,AGUUAA,AGUUAG,AGUUUA,AGUUUG,GGGGGG,GUAACG,UAGGAA,UAGGAU,UAGGUA,UAGGUU,UAGUAA,UAGUAU,UAGUUA,UAGUUU,UUUUUU |
| ZFP36L2 |
3 |
774 |
0.009523810 |
0.0039046755 |
1.286336 |
UAUUUA,UUAUUU |
AUUUAU,UAUUUA,UUAUUU,UUUAUU |
| A1CF |
2 |
598 |
0.007142857 |
0.0030179363 |
1.242939 |
UAAUUA,UUAAUU |
AGUAUA,AUAAUU,AUCAGU,CAGUAU,GAUCAG,UAAUUA,UAAUUG,UCAGUA,UGAUCA,UUAAUU |
| FMR1 |
26 |
5430 |
0.064285714 |
0.0273629585 |
1.232274 |
AAGGGA,AGCUGG,AGGAAU,AGGAUU,AGUGGC,AUGGAG,CUGAGG,GAGGAU,GCUGGG,GGCUGG,UAGCAG,UGAGGA,UGGAGU,UUUUUU |
AAAAAA,AAGCGG,AAGGAA,AAGGAG,AAGGAU,AAGGGA,ACUAAG,ACUGGG,ACUGGU,AGCAGC,AGCGAA,AGCGAC,AGCGGC,AGCUGG,AGGAAG,AGGAAU,AGGAGG,AGGAGU,AGGAUG,AGGAUU,AGGCAC,AGGGAG,AGUGGC,AUAGGC,AUGGAG,CAGCUG,CAUAGG,CGAAGG,CGACUG,CGGCUG,CUAAGG,CUAUGG,CUGAGG,CUGGGC,CUGGUG,GAAGGA,GAAGGG,GACAAG,GACAGG,GACUAA,GACUGG,GAGCGA,GAGCGG,GAGGAU,GCAGCU,GCAUAG,GCGAAG,GCGACU,GCGGCU,GCUAAG,GCUAUG,GCUGAG,GCUGGG,GGACAA,GGACAG,GGACUA,GGCAUA,GGCGAA,GGCUAA,GGCUAU,GGCUGA,GGCUGG,GGGCAU,GGGCGA,GGGCUA,GGGCUG,GGGGGG,GUGCGA,GUGCGG,GUGGCU,UAAGGA,UAGCAG,UAGCGA,UAGCGG,UAGGCA,UAGUGG,UAUGGA,UGACAA,UGACAG,UGAGGA,UGCGAC,UGCGGC,UGGAGU,UGGCUG,UUUUUU |
| HNRNPL |
6 |
1419 |
0.016666667 |
0.0071543732 |
1.220068 |
AAACAA,AACAAA,ACCACA,CACAAA,CACACC,CACCAC |
AAACAA,AAACAC,AAAUAA,AAAUAC,AACAAA,AACACA,AAUAAA,AAUACA,ACACAA,ACACAC,ACACCA,ACACGA,ACAUAA,ACAUAC,ACCACA,CACAAA,CACAAC,CACAAG,CACACA,CACACC,CACCAC,CACGAA,CACGAC,CACGAG,CAUAAA,CAUACA |
| PCBP2 |
19 |
4164 |
0.047619048 |
0.0209844821 |
1.182216 |
AAUUAC,AUUAAA,AUUACC,CCCUUA,CCUCCA,CCUCCU,CUCCCA,UUUUUU |
AAACCA,AAAUUA,AAAUUC,AACCAA,AACCCU,AAUUAA,AAUUAC,AAUUCA,AAUUCC,ACAUUA,ACCAAA,ACCCUA,ACCUUA,AUUAAA,AUUACC,AUUCAC,AUUCCA,AUUCCC,CAAUUA,CAUUAA,CCAUUA,CCAUUC,CCCCCA,CCCCCC,CCCCCU,CCCUAA,CCCUCC,CCCUUA,CCCUUC,CCUCCA,CCUCCC,CCUCCU,CCUUAA,CCUUCC,CUAACC,CUCCCA,CUCCCC,CUCCCU,CUUAAA,CUUCCA,CUUCCC,CUUCCU,GGGGGG,UAACCC,UCCCCA,UUAGAG,UUAGGA,UUAGGG,UUUUUU |
| ZNF638 |
2 |
646 |
0.007142857 |
0.0032597743 |
1.131729 |
UGUUCU,UGUUGU |
CGUUCG,CGUUCU,CGUUGG,CGUUGU,GGUUCG,GGUUCU,GGUUGG,GGUUGU,GUUCGU,GUUCUU,GUUGGU,GUUGUU,UGUUCG,UGUUCU,UGUUGG,UGUUGU |
| CELF2 |
3 |
881 |
0.009523810 |
0.0044437727 |
1.099754 |
GUGUGU,UGUGUG,UGUUGU |
AUGUGU,GUAUGU,GUCUGU,GUGUGU,GUUGUU,UAUGUG,UAUGUU,UGUGUG,UGUUGU,UUGUGU |
| PTBP1 |
33 |
8003 |
0.080952381 |
0.0403264813 |
1.005346 |
ACUUUC,ACUUUU,AUUUUC,AUUUUU,CUCCAU,CUUUUU,GUCUUU,UAGCUG,UAUUUU,UCCAUC,UCUUUU,UUAUUU,UUCUUU,UUUCUU,UUUUCU,UUUUUC,UUUUUU |
ACUUUC,ACUUUU,AGCUGU,AUCUUC,AUUUUC,AUUUUU,CAUCUU,CCAUCU,CCCCCC,CCUCUU,CCUUCC,CCUUUC,CCUUUU,CUAUCU,CUCCAU,CUCCCC,CUCUCU,CUCUUA,CUCUUC,CUCUUG,CUCUUU,CUUCCU,CUUCUC,CUUCUU,CUUUCC,CUUUCU,CUUUUC,CUUUUU,GCUCCC,GCUGUG,GGCUCC,GGGGGG,GUCUUA,GUCUUU,UACUUU,UAGCUG,UAUUUU,UCCAUC,UCCCCC,UCCUCU,UCUAUC,UCUCUC,UCUCUU,UCUUCU,UCUUUC,UCUUUU,UUACUU,UUAUUU,UUCCCC,UUCCUU,UUCUCU,UUCUUC,UUCUUG,UUCUUU,UUUCCC,UUUCUU,UUUUCC,UUUUCU,UUUUUC,UUUUUU |
| SFPQ |
29 |
7087 |
0.071428571 |
0.0357114067 |
1.000116 |
AAGAAC,AAGAGG,AAGGGA,AGAGGA,AGGGAU,CUGGAG,CUGGGA,GAGGAA,GGUAAG,GUAAGA,GUGAUU,UAAGAG,UAAGGG,UCUAAG,UGAUUU,UGGUUU,UUAAUU,UUUUUU |
AAGAAC,AAGAGC,AAGAGG,AAGCAA,AAGGAA,AAGGAC,AAGGGA,ACUGGG,AGAGAG,AGAGGA,AGAGGU,AGAUCG,AGGAAC,AGGACC,AGGGAU,AGGGGG,AUCGGA,CAGGCA,CUAAGG,CUGGAG,CUGGGA,GAAGAA,GAAGAG,GAAGCA,GAAGGA,GACUGG,GAGAGG,GAGGAA,GAGGAC,GAGGUA,GAUCGG,GCAGGC,GGAAGA,GGACUG,GGAGAG,GGAGGA,GGAGGG,GGGAUC,GGGGAC,GGGGAU,GGGGGA,GGGGGG,GGUAAG,GGUCUG,GUAAGA,GUAAUG,GUAAUU,GUAGUG,GUAGUU,GUCUGG,GUGAUG,GUGAUU,GUGGUG,GUGGUU,UAAGAG,UAAGGA,UAAGGG,UAAUGG,UAAUGU,UAAUUG,UAAUUU,UAGAGA,UAGAUC,UAGGGG,UAGUGG,UAGUGU,UAGUUG,UAGUUU,UCGGAA,UCUAAG,UCUGGA,UGAAGC,UGAUGG,UGAUGU,UGAUUG,UGAUUU,UGCAGG,UGGAAG,UGGAGA,UGGAGC,UGGAGG,UGGUCU,UGGUGG,UGGUGU,UGGUUG,UGGUUU,UUAAUG,UUAAUU,UUAGUG,UUAGUU,UUGAAG,UUGAUG,UUGAUU,UUGGUG,UUGGUU,UUUUUU |